Summary information and primary citation
- PDB-id
- 7luf; DSSR-derived features in text and JSON formats
- Class
- transferase-DNA
- Method
- X-ray (3.5 Å)
- Summary
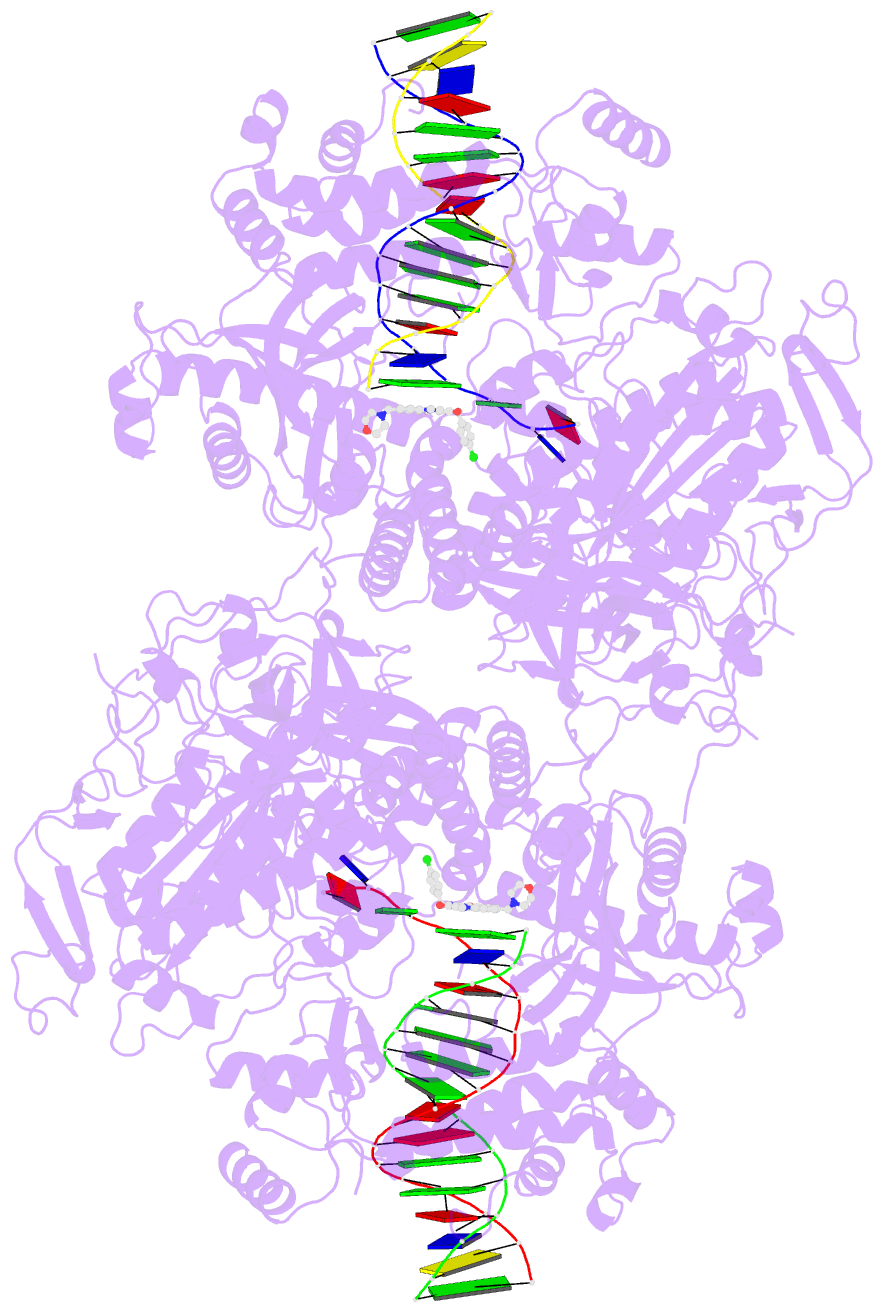
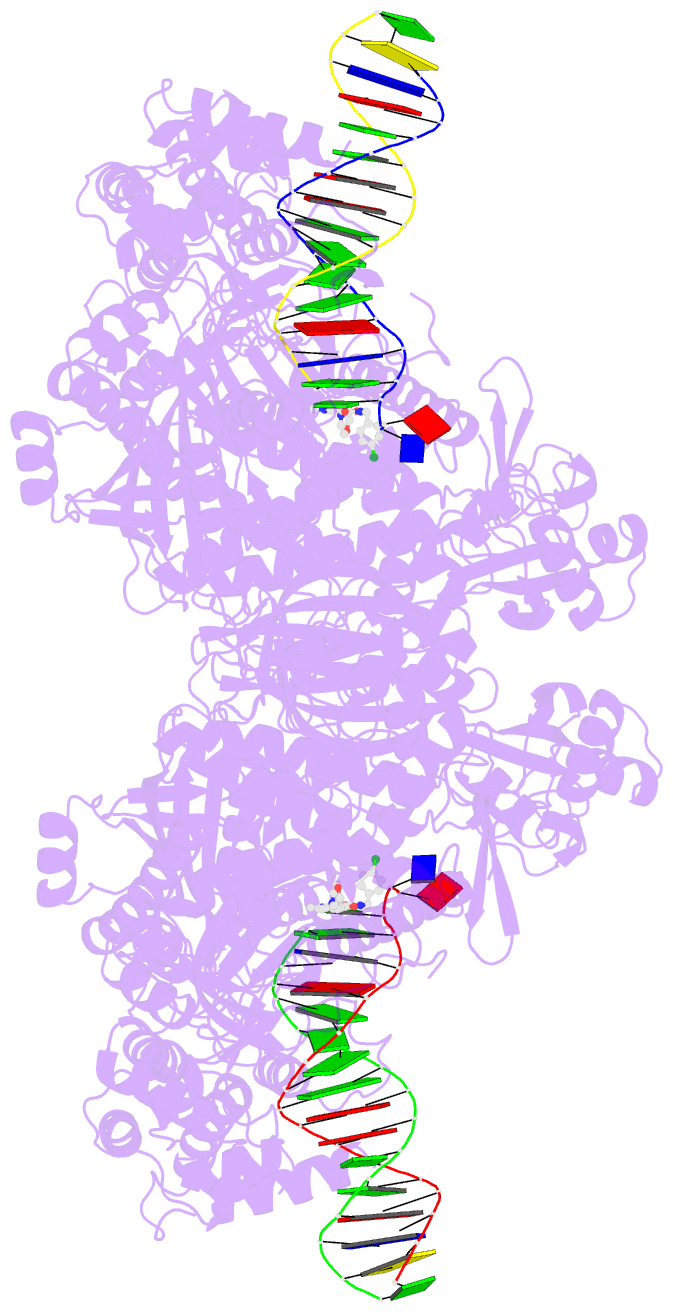
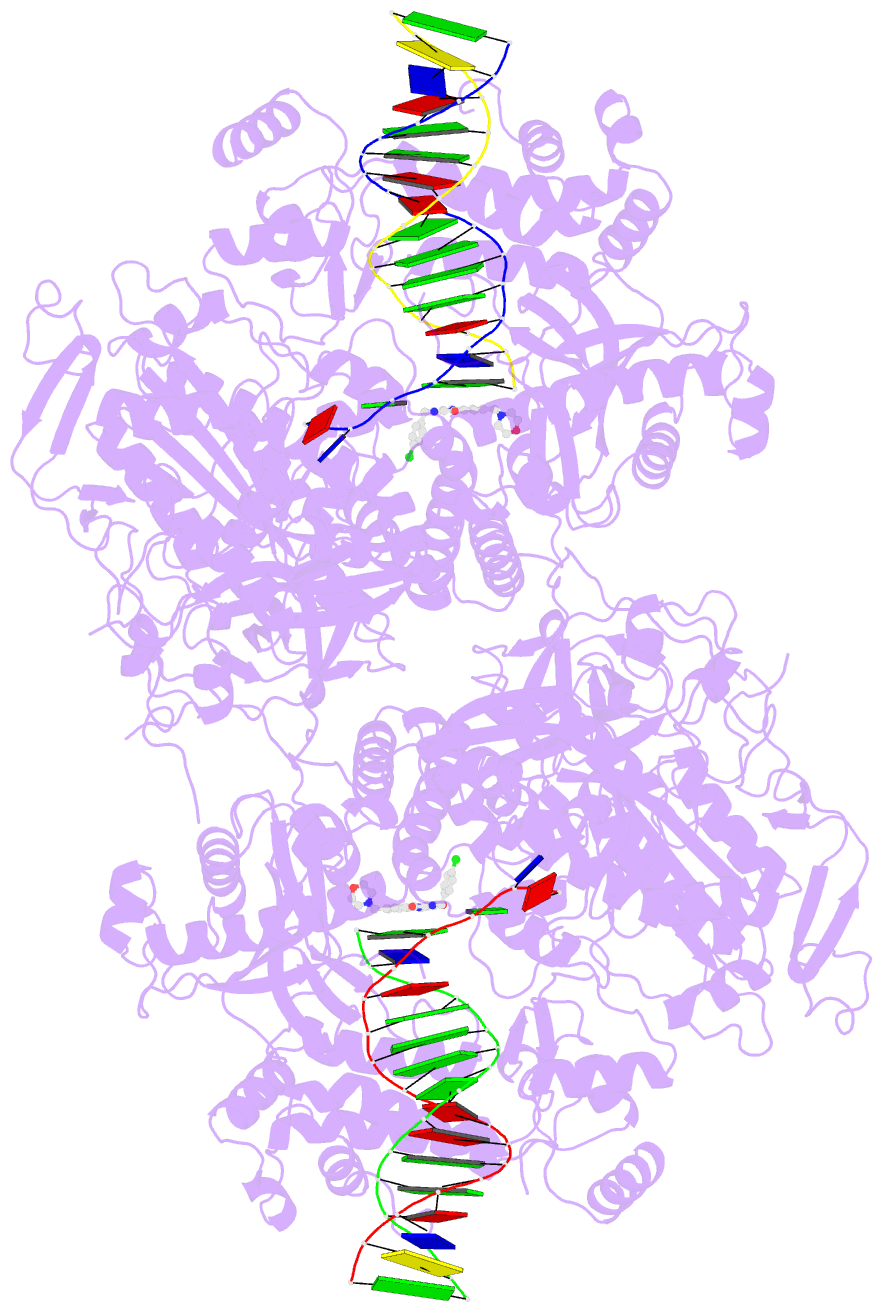
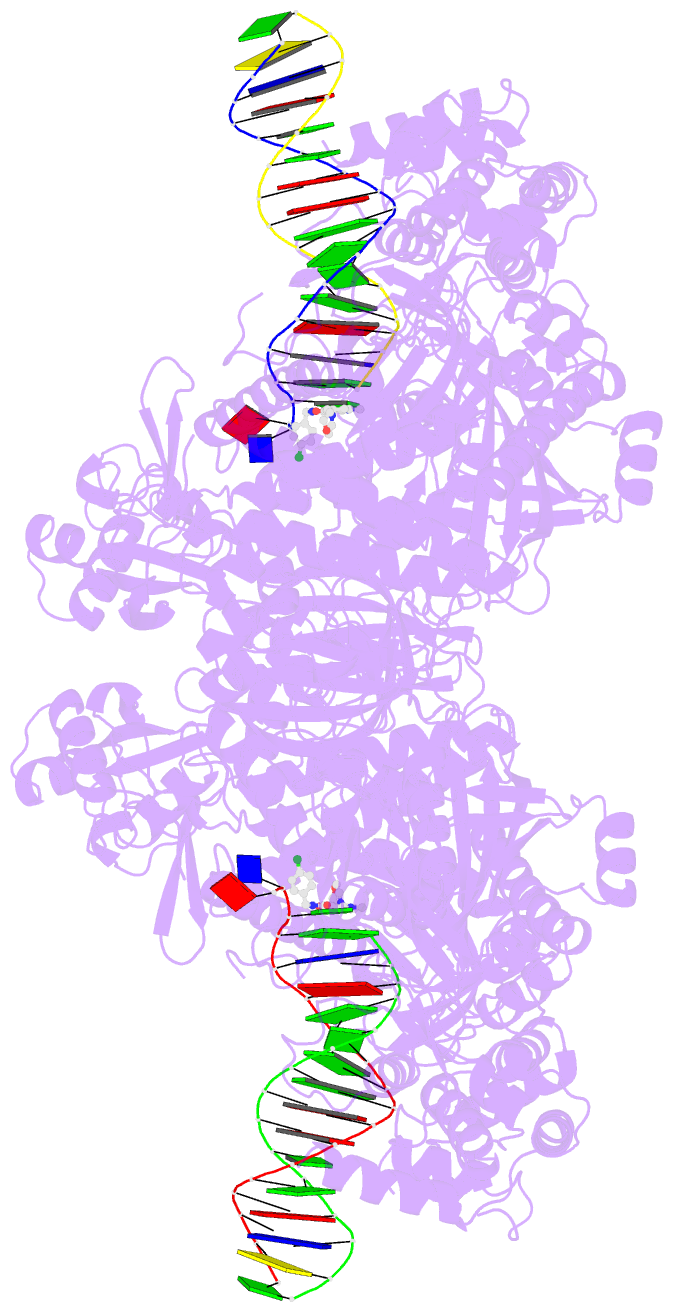
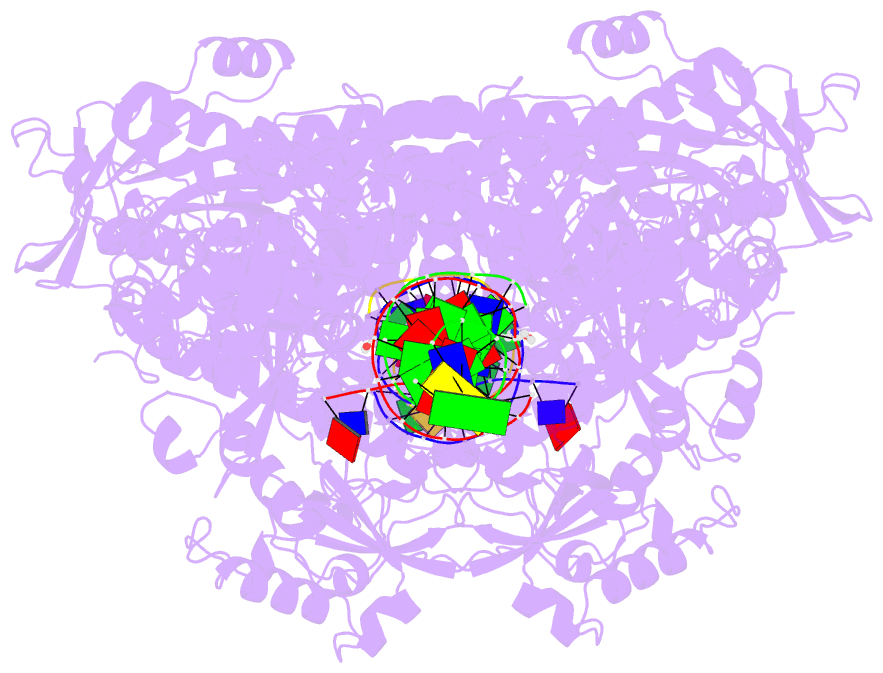
- Hsv1 polymerase ternary complex with dsDNA and pnu-183792
- Reference
- Hayes RP, Heo MR, Mason M, Reid J, Burlein C, Armacost KA, Tellers DM, Raheem I, Shaw AW, Murray E, McKenna PM, Abeywickrema P, Sharma S, Soisson SM, Klein D (2021): "Structural understanding of non-nucleoside inhibition in an elongating herpesvirus polymerase." Nat Commun, 12, 3040. doi: 10.1038/s41467-021-23312-8.
- Abstract
- All herpesviruses encode a conserved DNA polymerase that is required for viral genome replication and serves as an important therapeutic target. Currently available herpesvirus therapies include nucleoside and non-nucleoside inhibitors (NNI) that target the DNA-bound state of herpesvirus polymerase and block replication. Here we report the ternary complex crystal structure of Herpes Simplex Virus 1 DNA polymerase bound to DNA and a 4-oxo-dihydroquinoline NNI, PNU-183792 (PNU), at 3.5 Å resolution. PNU bound at the polymerase active site, displacing the template strand and inducing a conformational shift of the fingers domain into an open state. These results demonstrate that PNU inhibits replication by blocking association of dNTP and stalling the enzyme in a catalytically incompetent conformation, ultimately acting as a nucleotide competing inhibitor (NCI). Sequence conservation of the NCI binding pocket further explains broad-spectrum activity while a direct interaction between PNU and residue V823 rationalizes why mutations at this position result in loss of inhibition.