Summary information and primary citation
- PDB-id
- 7c9c; DSSR-derived features in text and JSON formats
- Class
- recombination-DNA
- Method
- cryo-EM (3.33 Å)
- Summary
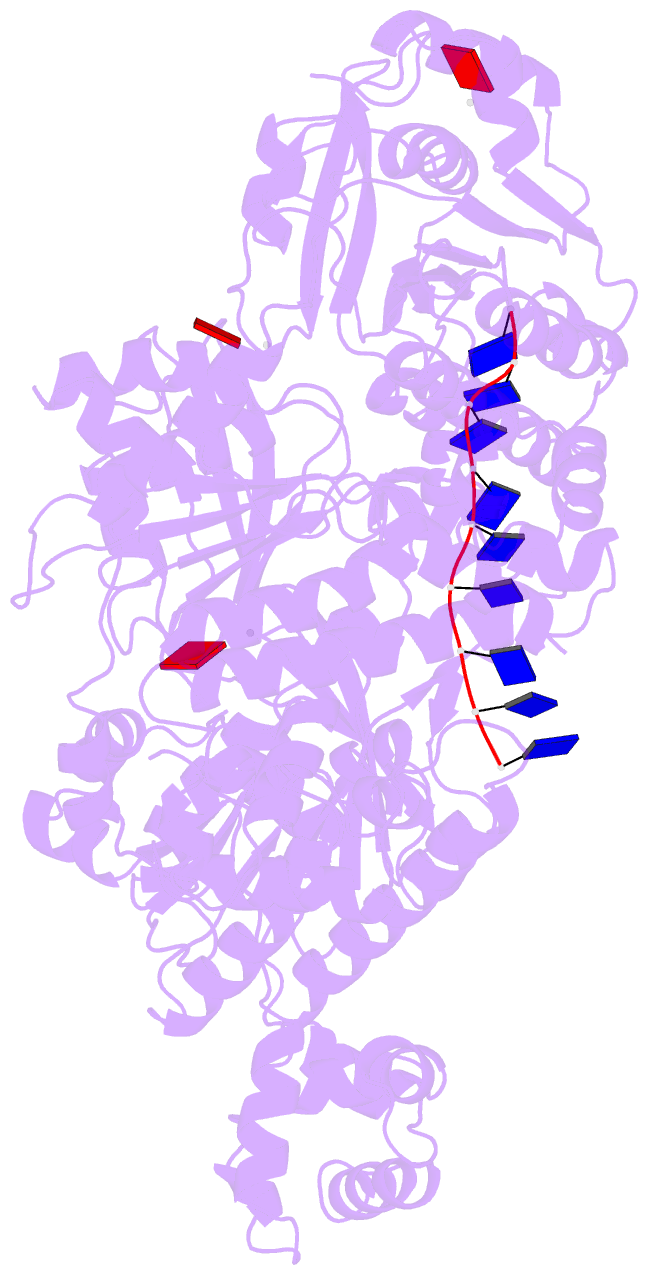
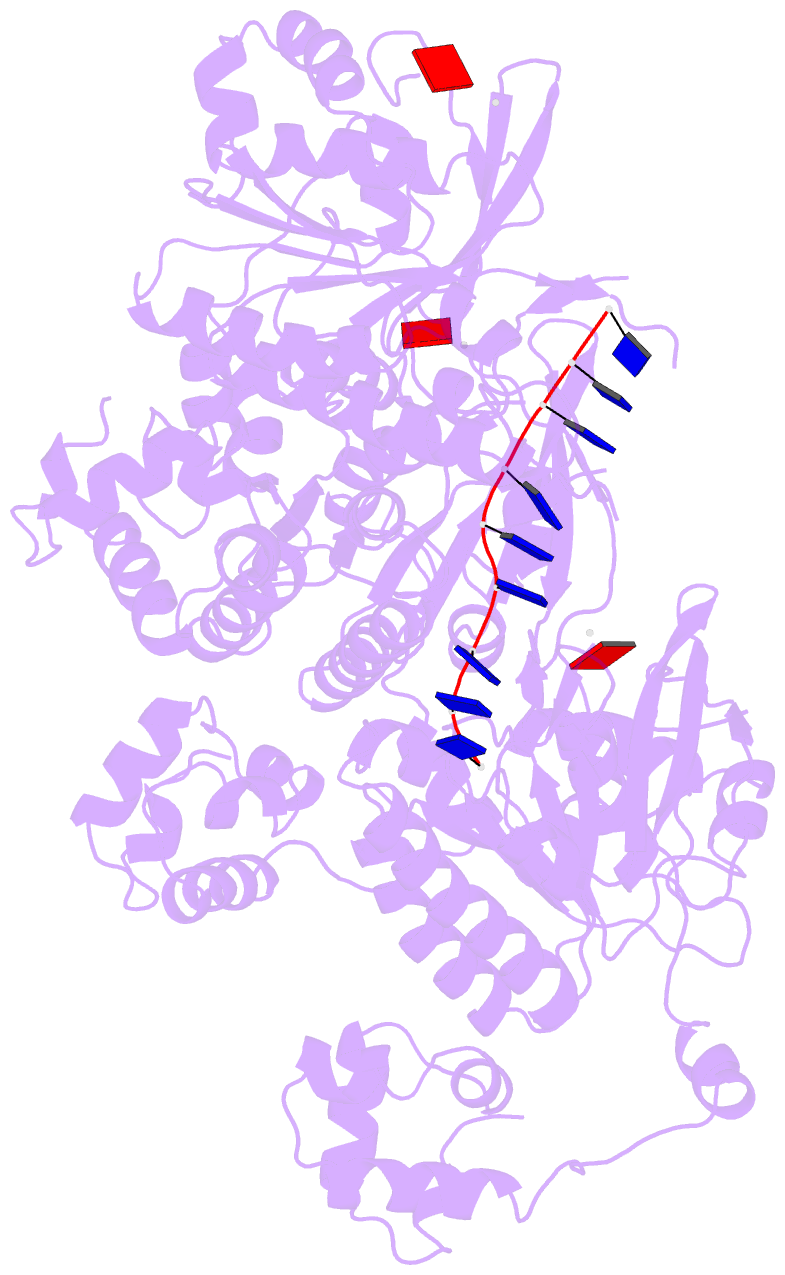
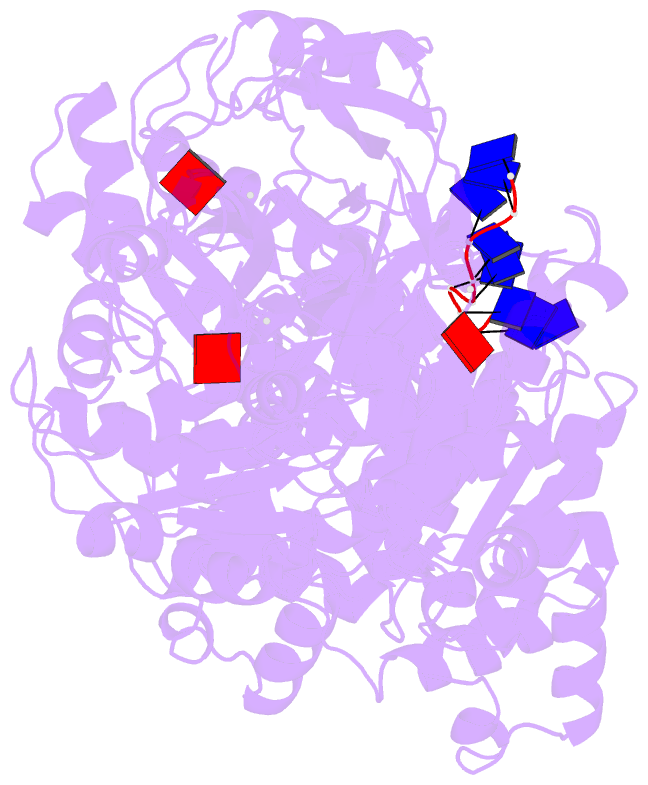
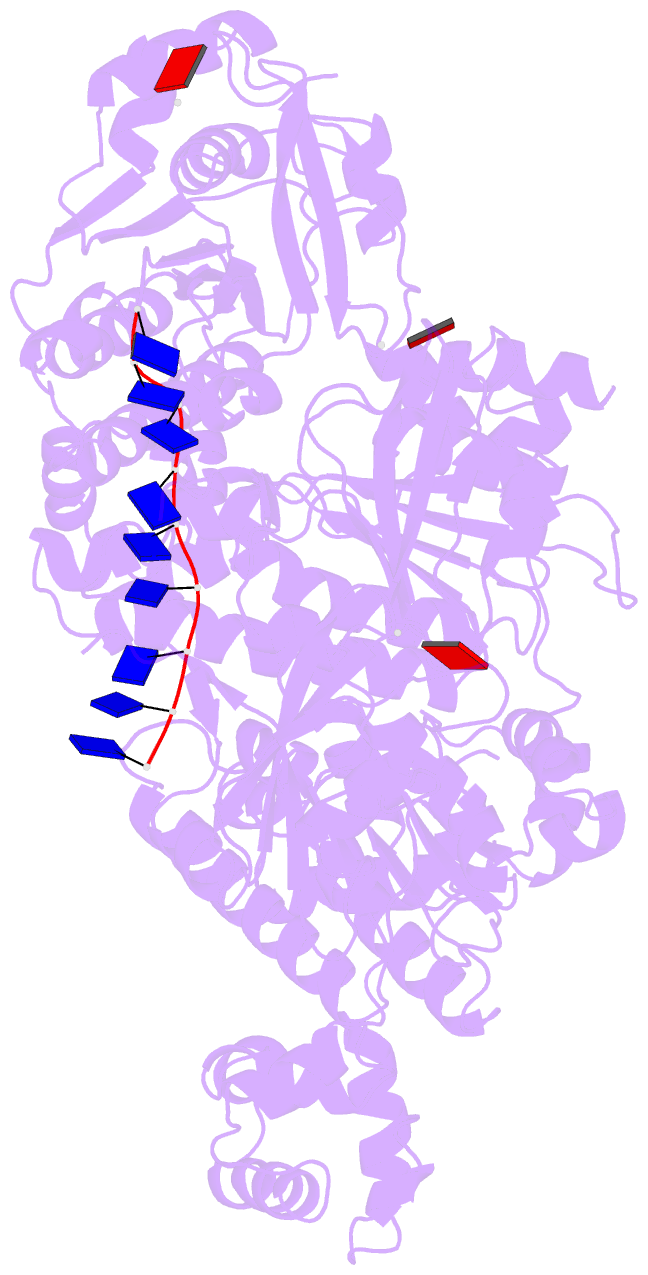
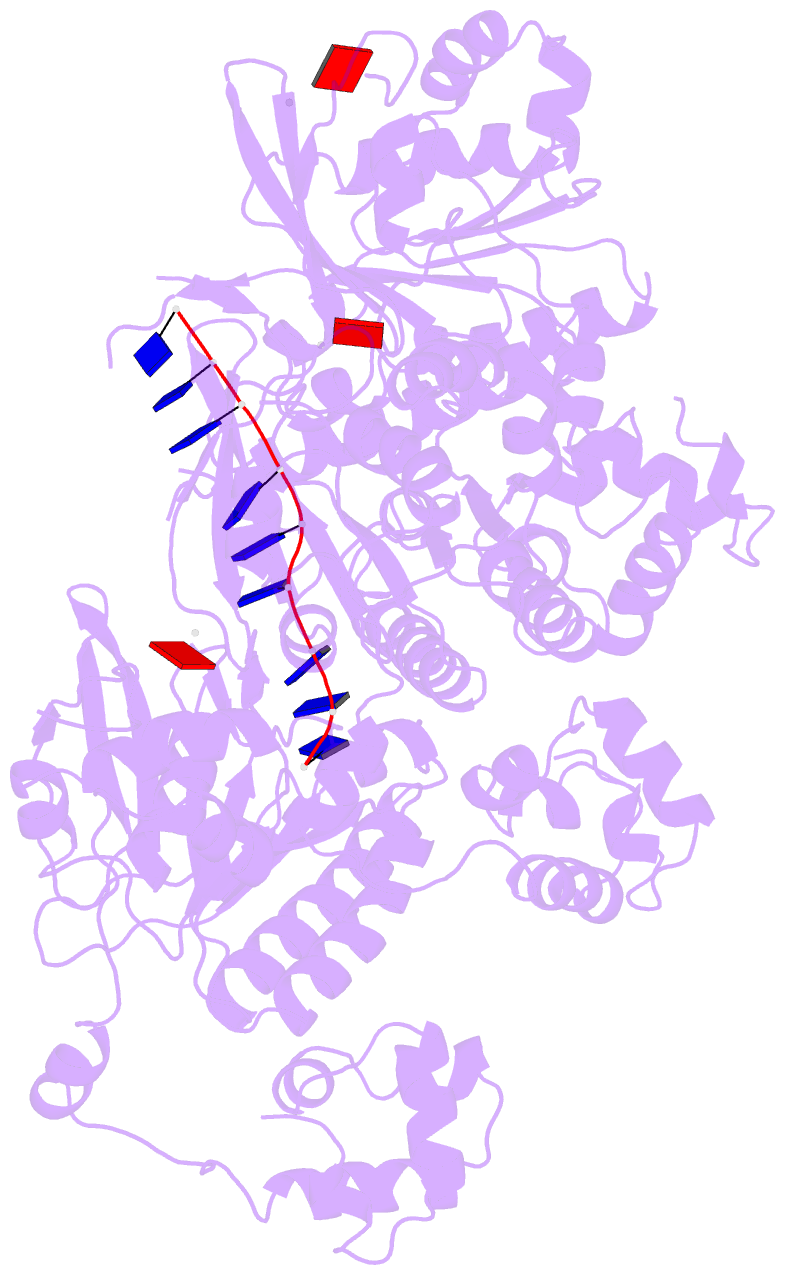
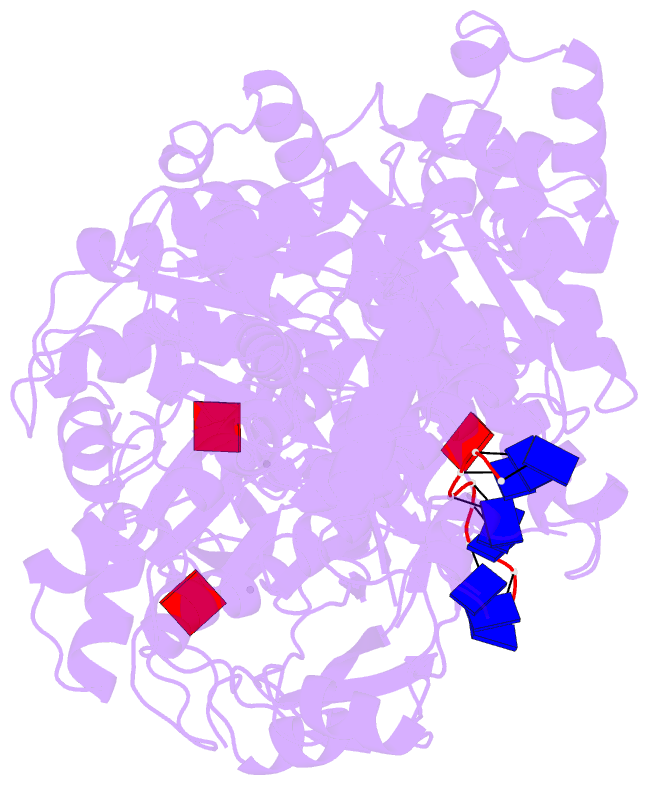
- Human dmc1 pre-synaptic complexes
- Reference
- Luo SC, Yeh HY, Lan WH, Wu YM, Yang CH, Chang HY, Su GC, Lee CY, Wu WJ, Li HW, Ho MC, Chi P, Tsai MD (2021): "Identification of fidelity-governing factors in human recombinases DMC1 and RAD51 from cryo-EM structures." Nat Commun, 12, 115. doi: 10.1038/s41467-020-20258-1.
- Abstract
- Both high-fidelity and mismatch-tolerant recombination, catalyzed by RAD51 and DMC1 recombinases, respectively, are indispensable for genomic integrity. Here, we use cryo-EM, MD simulation and functional analysis to elucidate the structural basis for the mismatch tolerance of DMC1. Structural analysis of DMC1 presynaptic and postsynaptic complexes suggested that the lineage-specific Loop 1 Gln244 (Met243 in RAD51) may help stabilize DNA backbone, whereas Loop 2 Pro274 and Gly275 (Val273/Asp274 in RAD51) may provide an open "triplet gate" for mismatch tolerance. In support, DMC1-Q244M displayed marked increase in DNA dynamics, leading to unobservable DNA map. MD simulation showed highly dispersive mismatched DNA ensemble in RAD51 but well-converged DNA in DMC1 and RAD51-V273P/D274G. Replacing Loop 1 or Loop 2 residues in DMC1 with RAD51 counterparts enhanced DMC1 fidelity, while reciprocal mutations in RAD51 attenuated its fidelity. Our results show that three Loop 1/Loop 2 residues jointly enact contrasting fidelities of DNA recombinases.