Summary information and primary citation
- PDB-id
-
6rjg;
SNAP-derived features in text and
JSON formats
- Class
- hydrolase
- Method
- cryo-EM (3.2 Å)
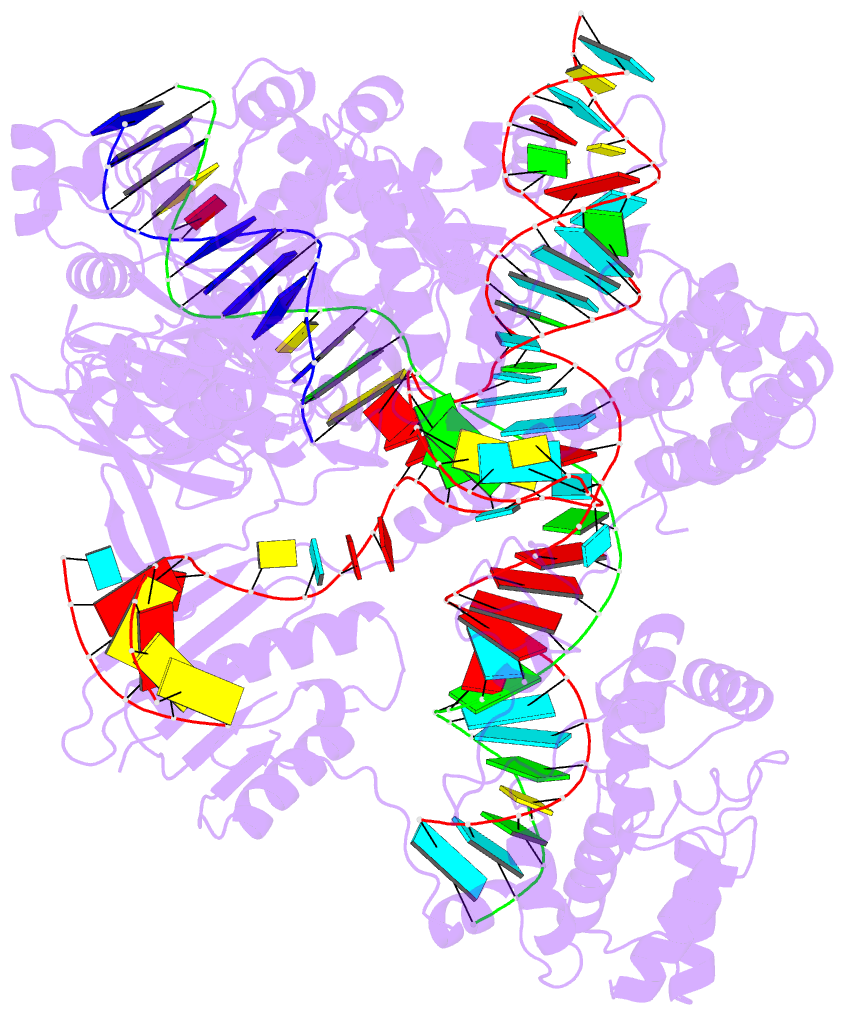
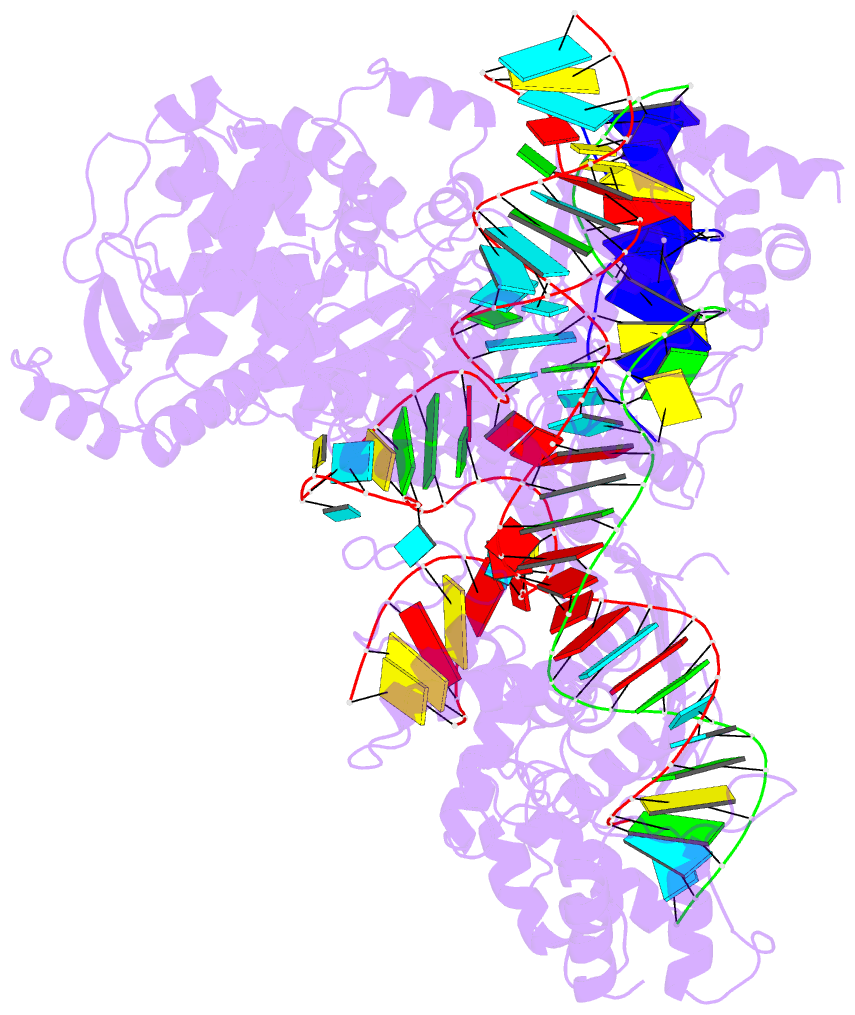
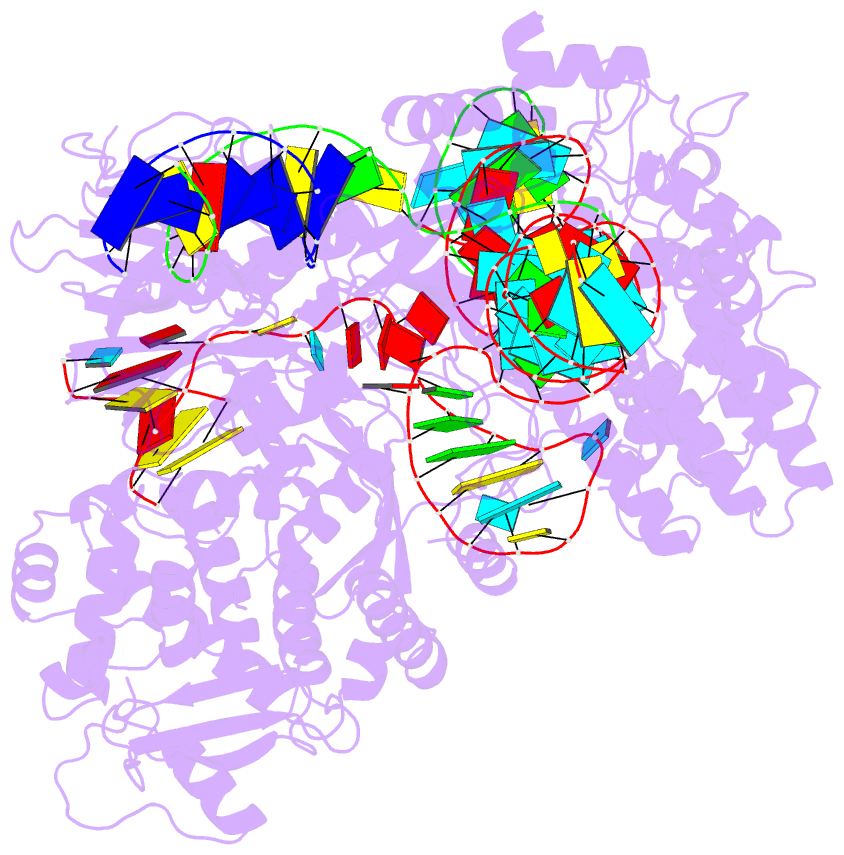
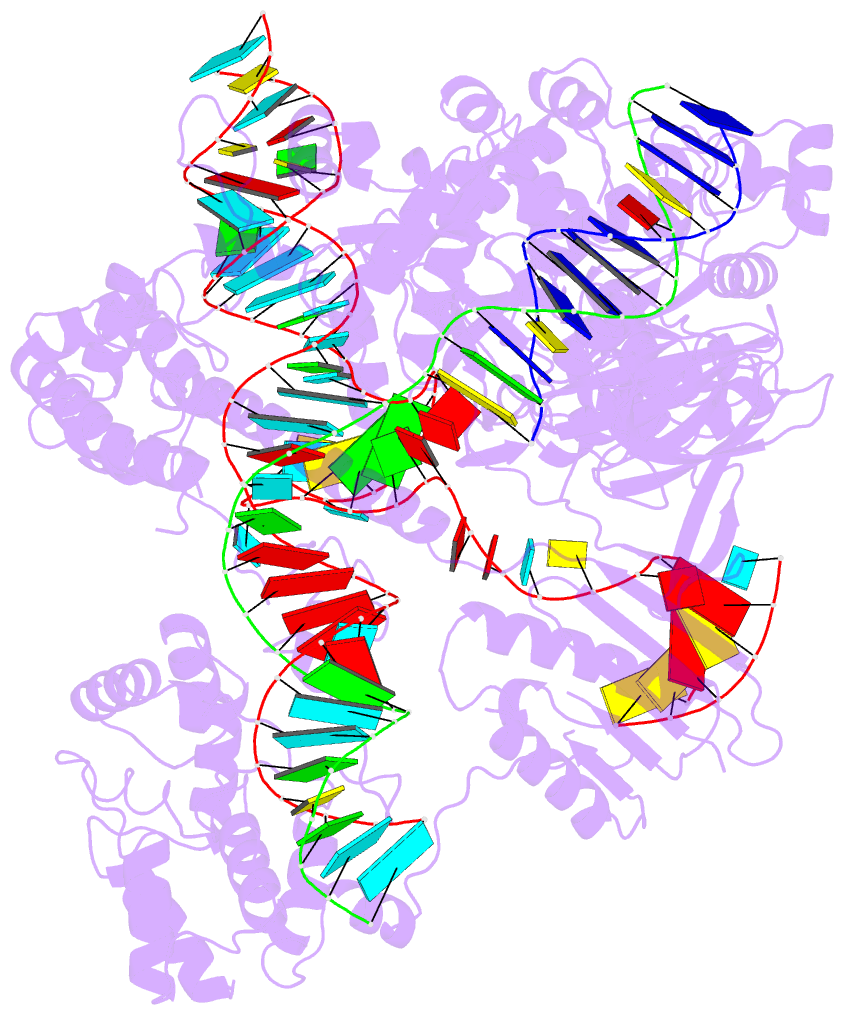
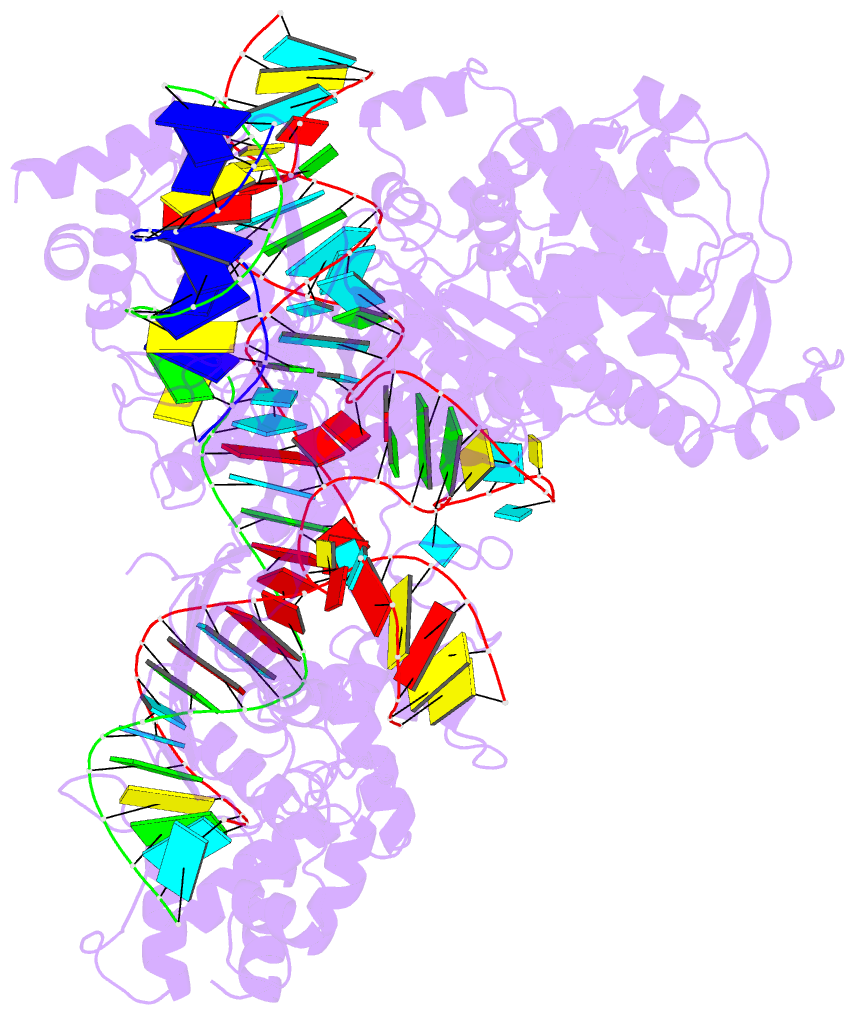
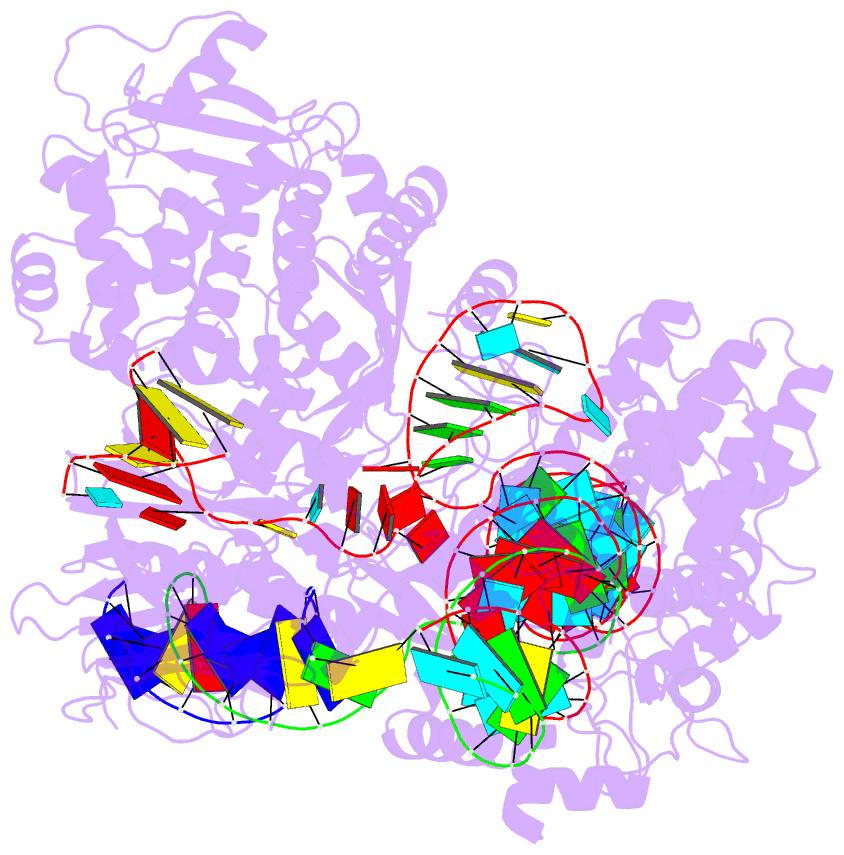
- Summary
- cryo-EM structure of st1cas9-sgrna-acriia6-tdna59-ntpam
complex.
- Reference
-
Fuchsbauer O, Swuec P, Zimberger C, Amigues B, Levesque
S, Agudelo D, Duringer A, Chaves-Sanjuan A, Spinelli S,
Rousseau GM, Velimirovic M, Bolognesi M, Roussel A,
Cambillau C, Moineau S, Doyon Y, Goulet A (2019):
"Cas9
Allosteric Inhibition by the Anti-CRISPR Protein
AcrIIA6." Mol.Cell, 76,
922. doi: 10.1016/j.molcel.2019.09.012.
- Abstract
- In the arms race against bacteria, bacteriophages have
evolved diverse anti-CRISPR proteins (Acrs) that block
CRISPR-Cas immunity. Acrs play key roles in the molecular
coevolution of bacteria with their predators, use a variety
of mechanisms of action, and provide tools to regulate
Cas-based genome manipulation. Here, we present structural
and functional analyses of AcrIIA6, an Acr from virulent
phages, exploring its unique anti-CRISPR action. Our
cryo-EM structures and functional data of AcrIIA6 binding
to Streptococcus thermophilus Cas9 (St1Cas9) show that
AcrIIA6 acts as an allosteric inhibitor and induces St1Cas9
dimerization. AcrIIA6 reduces St1Cas9 binding affinity for
DNA and prevents DNA binding within cells. The PAM and
AcrIIA6 recognition sites are structurally close and
allosterically linked. Mechanistically, AcrIIA6 affects the
St1Cas9 conformational dynamics associated with PAM
binding. Finally, we identify a natural St1Cas9 variant
resistant to AcrIIA6 illustrating Acr-driven mutational
escape and molecular diversification of Cas9 proteins.