Summary information and primary citation
- PDB-id
-
6ldm;
DSSR-derived features in text and
JSON formats
- Class
- DNA binding protein-DNA
- Method
- X-ray (2.4 Å)
- Summary
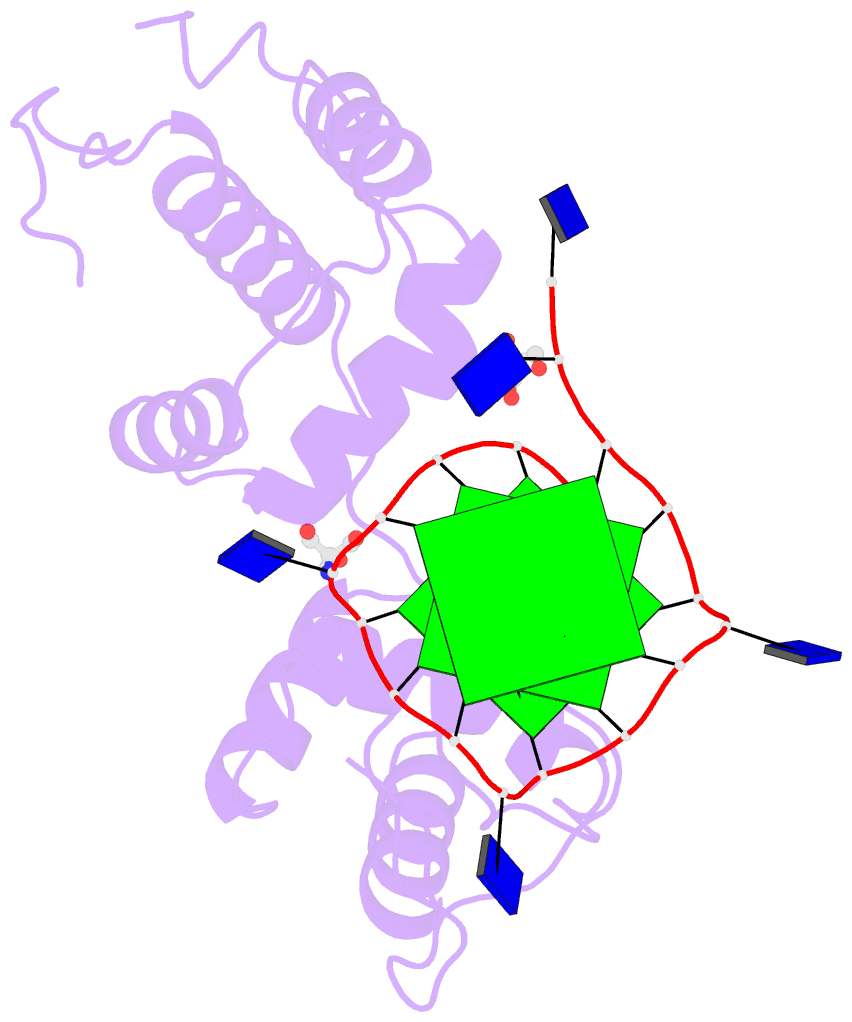
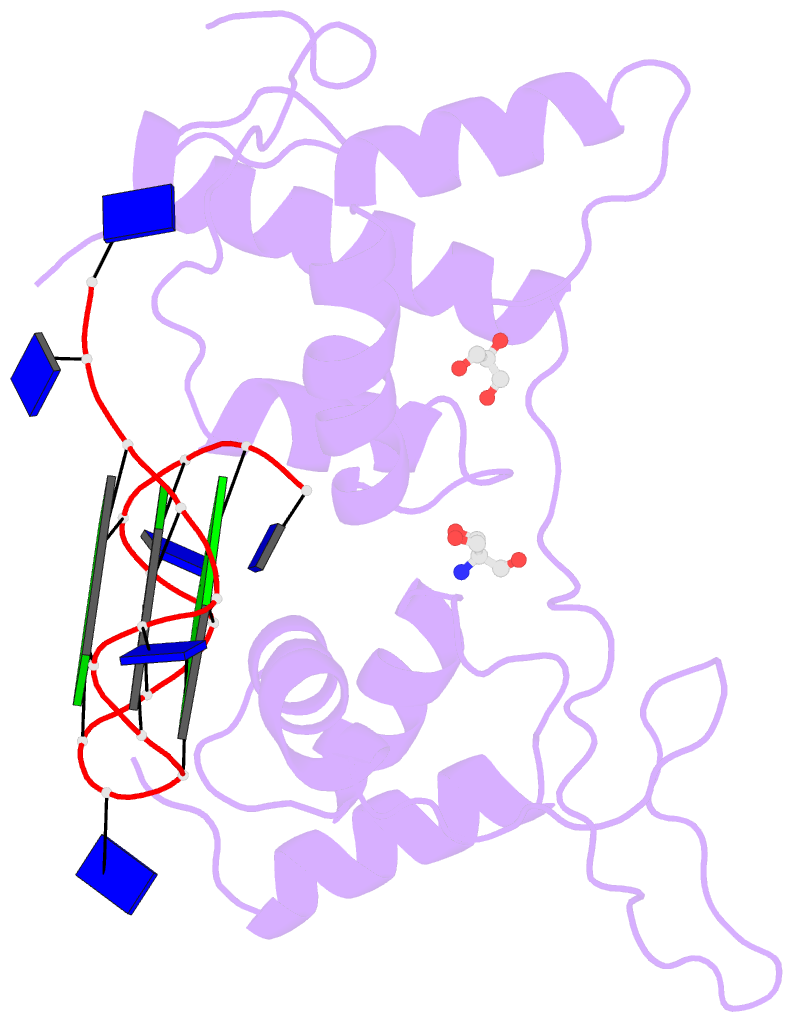
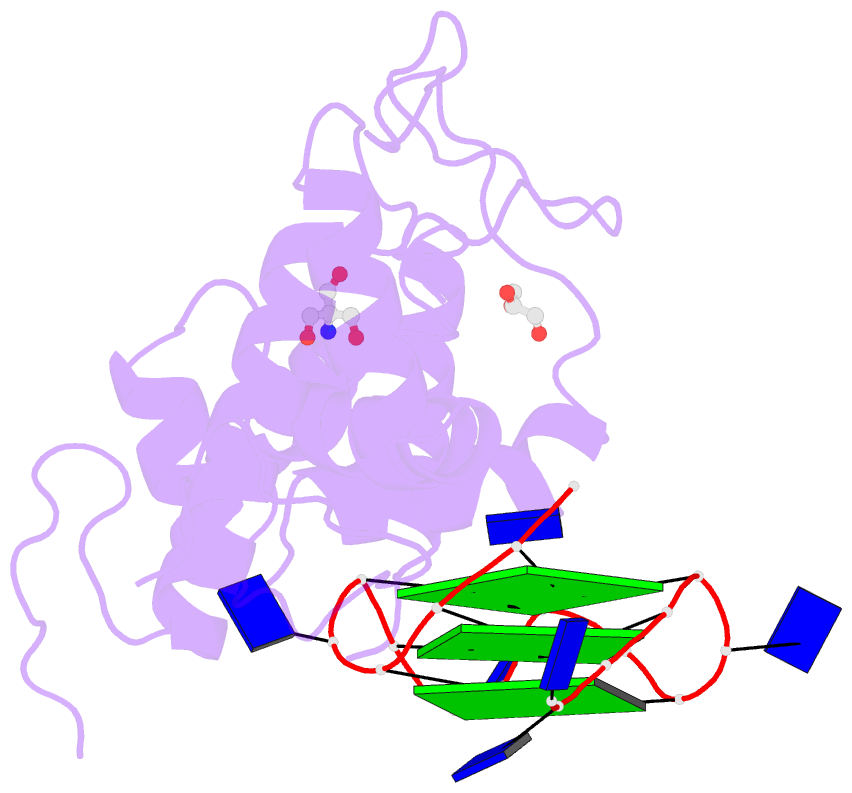
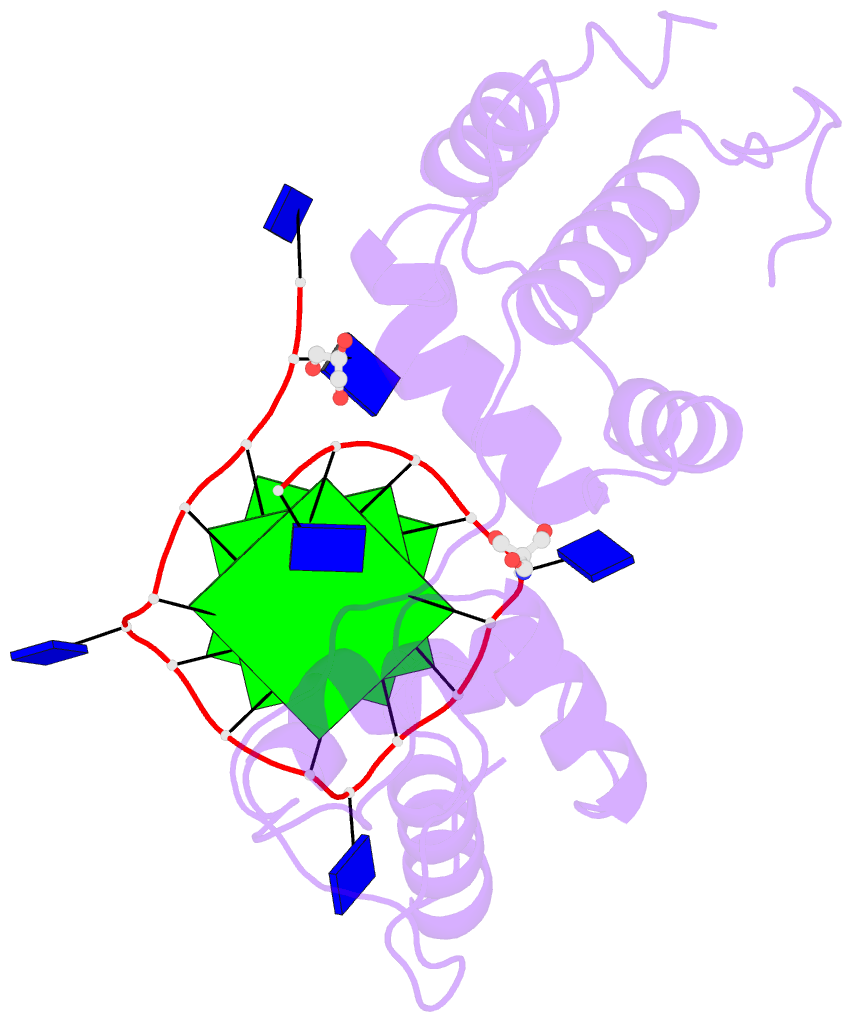
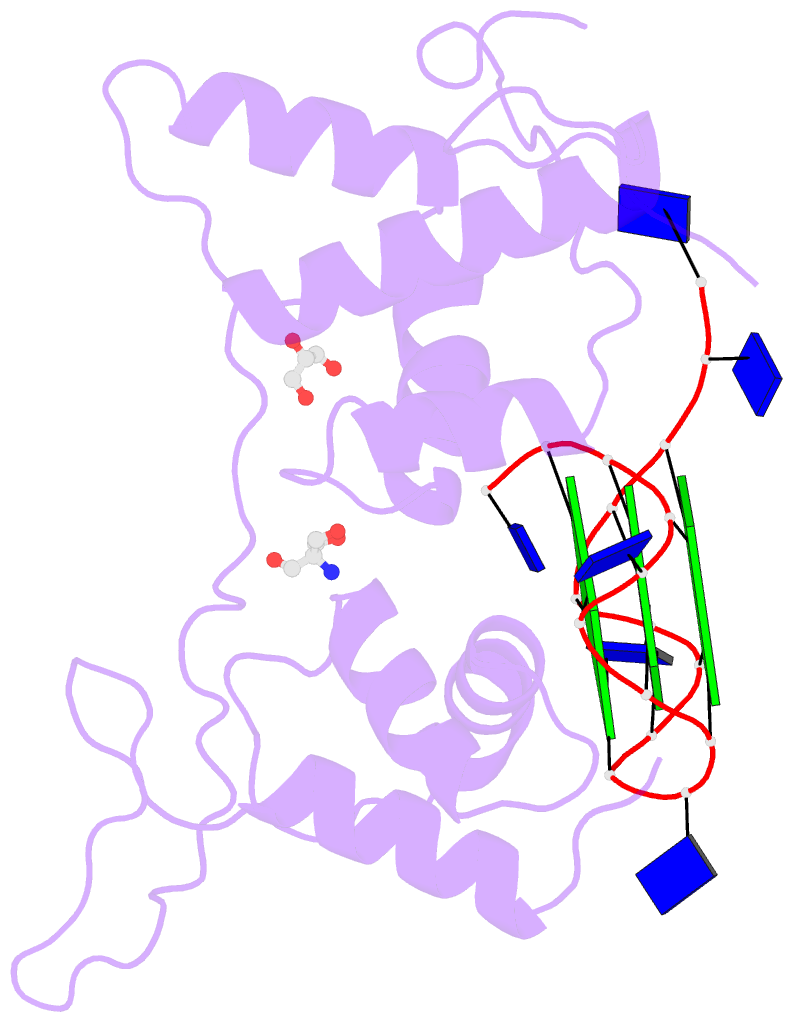
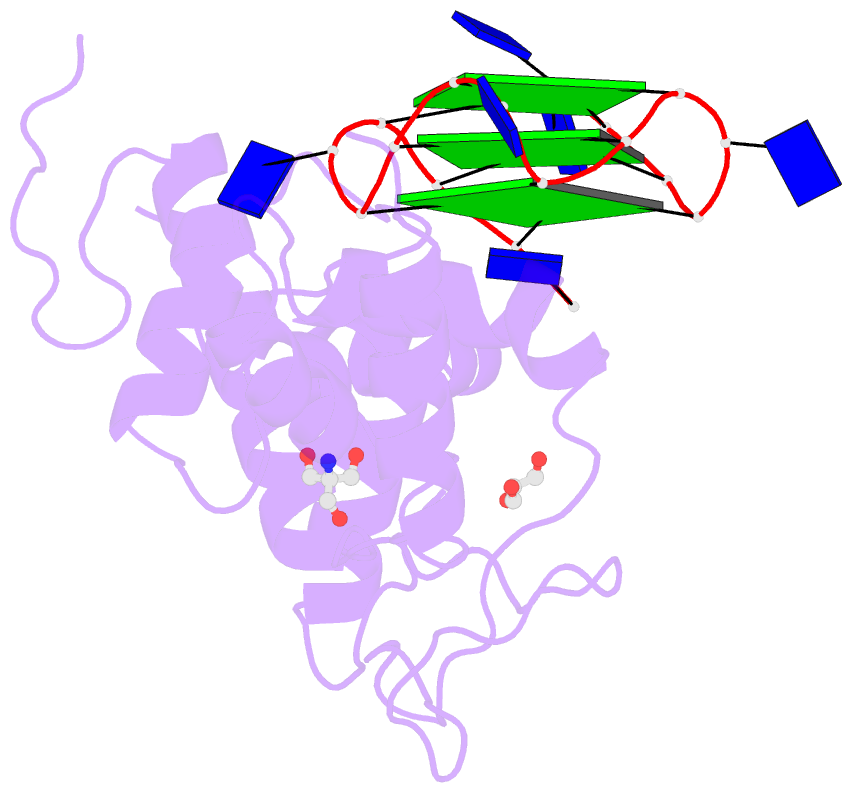
- Structural basis of g-quadruplex DNA recognition by the
yeast telomeric protein rap1
- Reference
-
Traczyk A, Liew CW, Gill DJ, Rhodes D (2020): "Structural
basis of G-quadruplex DNA recognition by the yeast
telomeric protein Rap1." Nucleic Acids Res.,
48, 4562-4571. doi: 10.1093/nar/gkaa171.
- Abstract
- G-quadruplexes are four-stranded nucleic acid
structures involved in multiple cellular pathways including
DNA replication and telomere maintenance. Such structures
are formed by G-rich DNA sequences typified by telomeric
DNA repeats. Whilst there is evidence for proteins that
bind and regulate G-quadruplex formation, the molecular
basis for this remains poorly understood. The budding yeast
telomeric protein Rap1, originally identified as a
transcriptional regulator functioning by recognizing
double-stranded DNA binding sites, was one of the first
proteins to be discovered to also bind and promote
G-quadruplex formation in vitro. Here, we present the 2.4 Å
resolution crystal structure of the Rap1 DNA-binding domain
in complex with a G-quadruplex. Our structure not only
provides a detailed insight into the structural basis for
G-quadruplex recognition by a protein, but also gives
a mechanistic understanding of how the same DNA-binding
domain adapts to specifically recognize different DNA
structures. The key observation is the DNA-recognition
helix functions in a bimodal manner: In double-stranded DNA
recognition one helix face makes electrostatic interactions
with the major groove of DNA, whereas in G-quadruplex
recognition a different helix face is used to make
primarily hydrophobic interactions with the planar face of
a G-tetrad.