Summary information and primary citation
- PDB-id
- 4fjk; DSSR-derived features in text and JSON formats
- Class
- transferase-DNA
- Method
- X-ray (2.0 Å)
- Summary
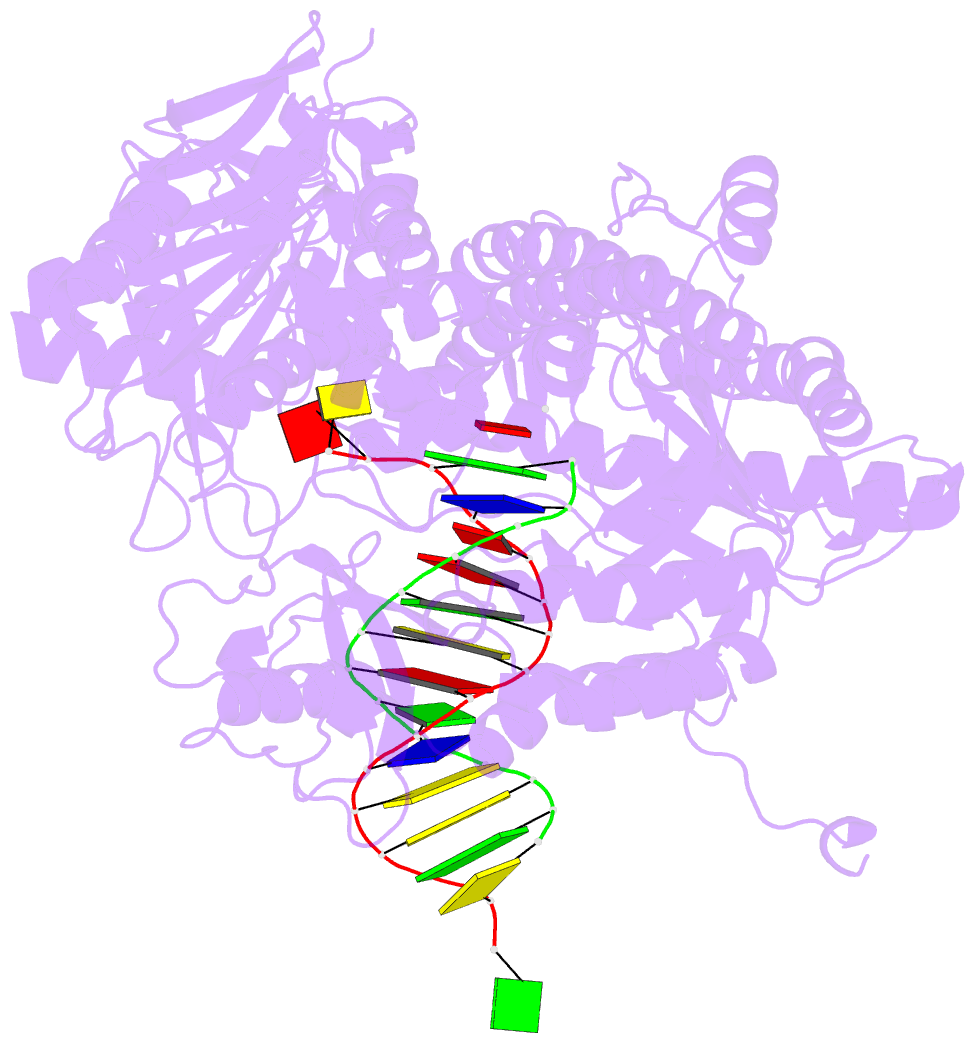
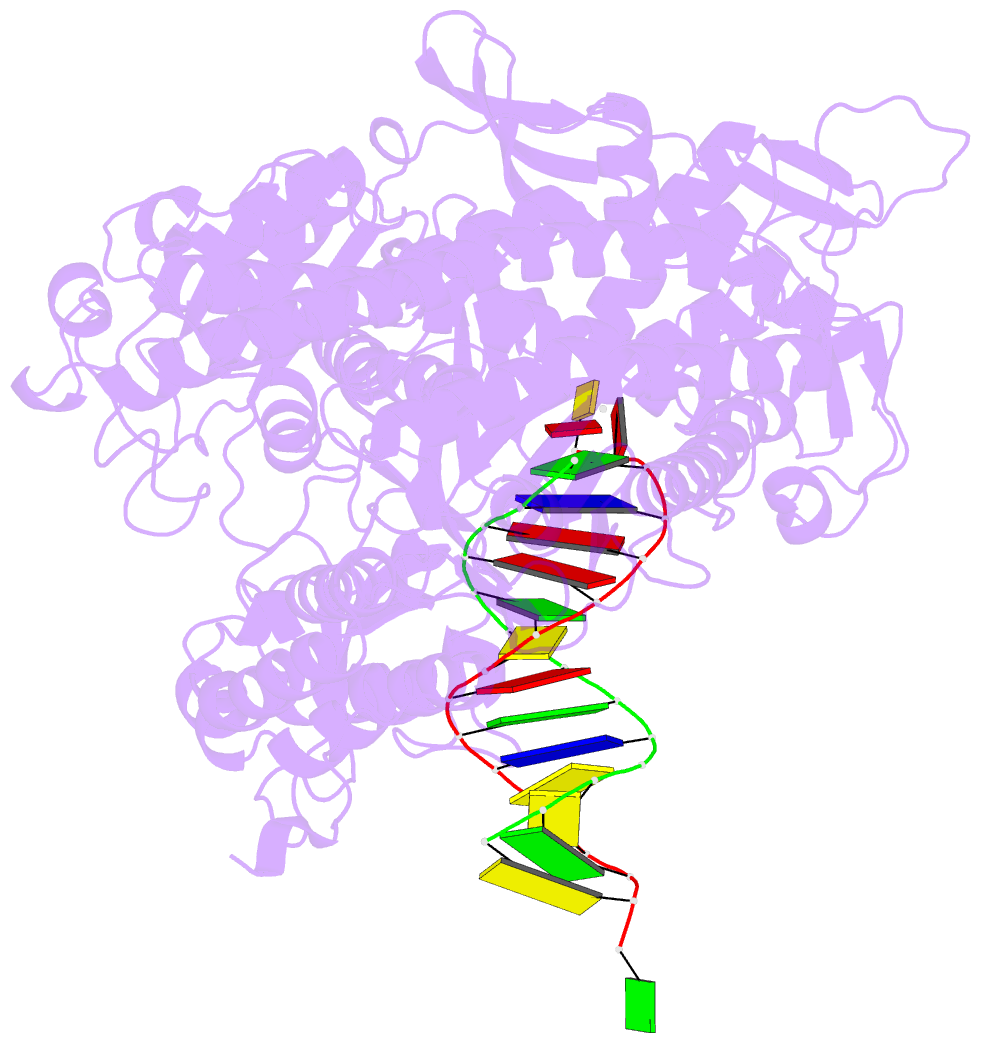
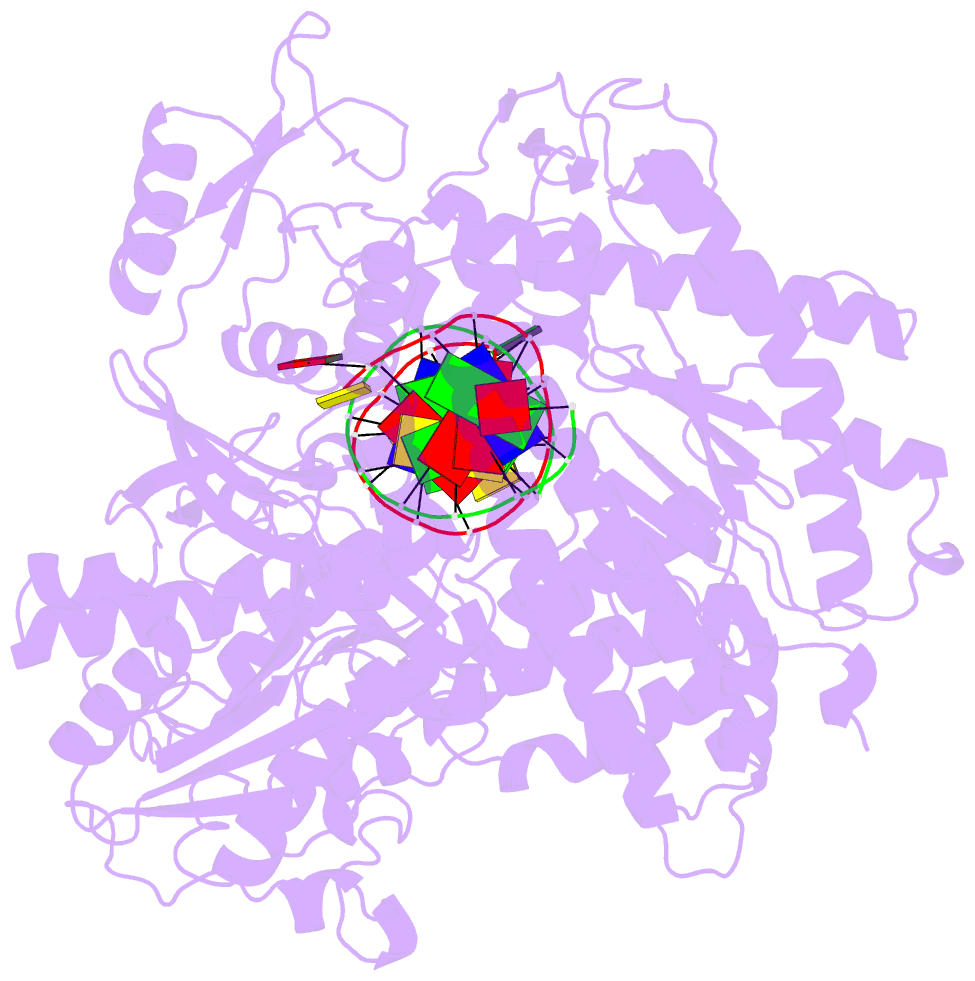
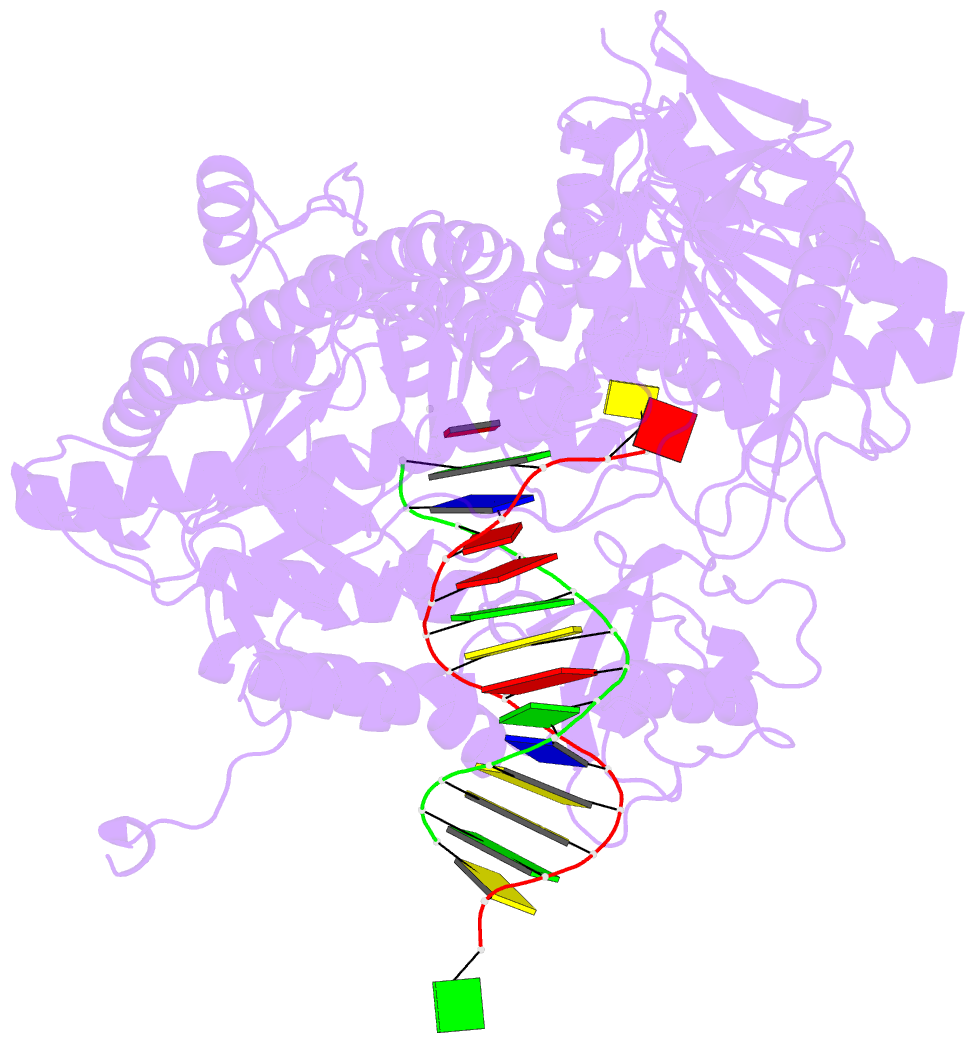
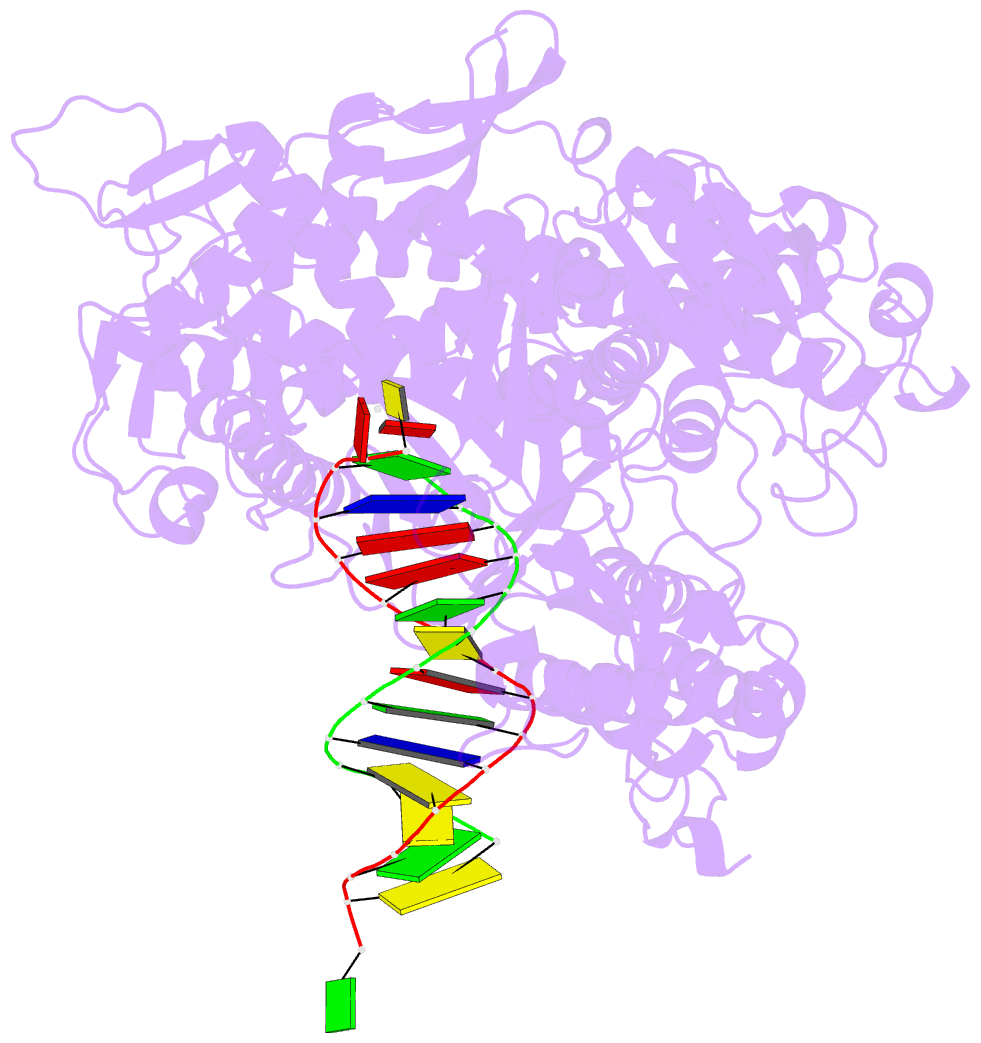
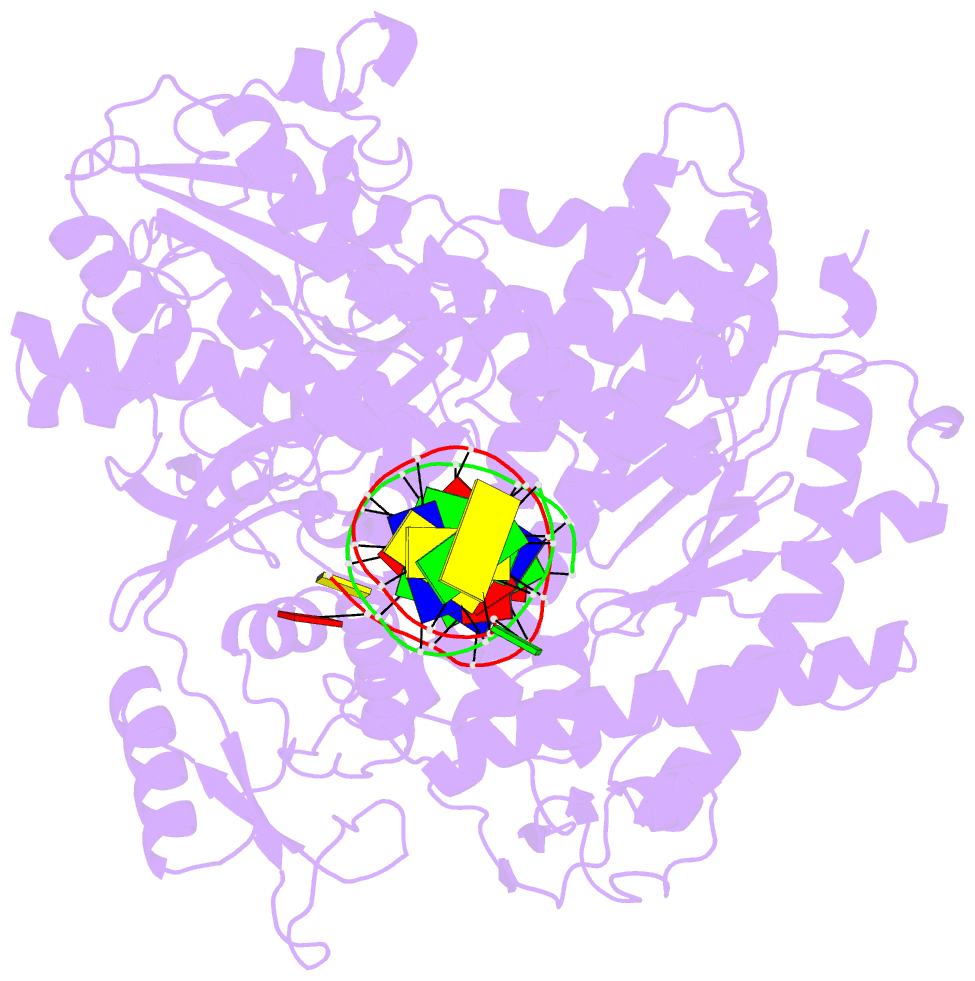
- Rb69 DNA polymerase ternary complex with datp-da
- Reference
- Xia S, Wang J, Konigsberg WH (2013): "DNA mismatch synthesis complexes provide insights into base selectivity of a B family DNA polymerase." J.Am.Chem.Soc., 135, 193-202. doi: 10.1021/ja3079048.
- Abstract
- Current hypotheses that attempt to rationalize the high degree of base selectivity exhibited by replicative DNA polymerases (pols) concur that ternary complexes formed with incorrect dNTPs are destabilized. Knowing what accounts for this destabilization is likely to be the key to understanding base discrimination. To address this issue, we have determined crystal structures of ternary complexes with all twelve mismatches using an engineered RB69 pol quadruple mutant (qm, L415A/L561A/S565G/Y567A) that enabled us to capture nascent mispaired dNTPs. These structures show that mismatches in the Nascent base-pair Binding Pocket (NBP) of the qm pol differ markedly from mismatches embedded in binary pol-DNA complexes. Surprisingly only 3 of 12 mismatches clash with the NBP when they are modeled into the wt pol. The remaining can fit into a wt pol ternary complex but deviate from normal Watson-Crick base-pairs. Repositioning of the templating nucleotide residue and the enlarged NBP in qm ternary complex play important roles in accommodating incorrect incoming dNTPs. From these structures, we propose additional reasons as to why incorrect dNTP are incorporated so inefficiently by wild type RB69 pol: (i) steric clashes with side chains in the NBP after Fingers closing; (ii) weak interactions or large gaps between the incoming dNTP and the templating base; (iii) burying a protonated base in the hydrophobic environment of the NBP. All of these possibilities would be expected to destabilize the closed ternary complex so that incorporation of incorrect dNTP would be a rare event.