Summary information and primary citation
- PDB-id
- 3n1l; DSSR-derived features in text and JSON formats
- Class
- DNA binding protein-DNA
- Method
- X-ray (2.35 Å)
- Summary
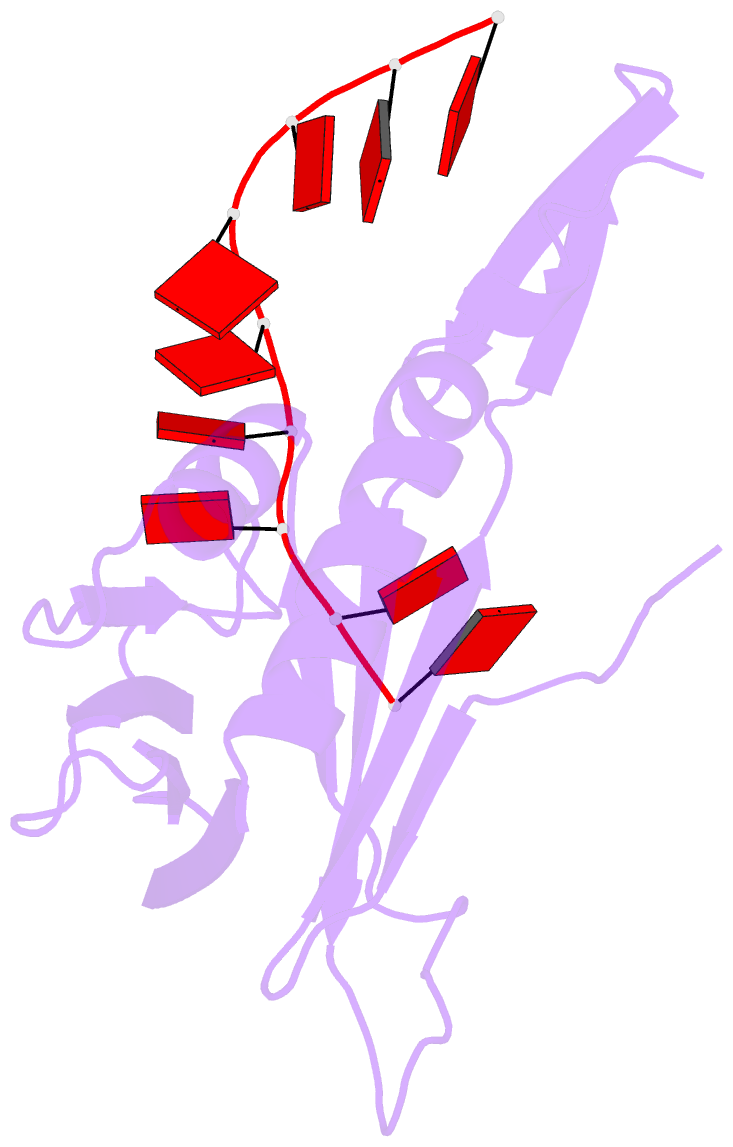
- Crystal structure of a stwhy2-rcere32 complex
- Reference
- Cappadocia L, Marechal A, Parent JS, Lepage E, Sygusch J, Brisson N (2010): "Crystal Structures of DNA-Whirly Complexes and Their Role in Arabidopsis Organelle Genome Repair." Plant Cell, 22, 1849-1867. doi: 10.1105/tpc.109.071399.
- Abstract
- DNA double-strand breaks are highly detrimental to all organisms and need to be quickly and accurately repaired. Although several proteins are known to maintain plastid and mitochondrial genome stability in plants, little is known about the mechanisms of DNA repair in these organelles and the roles of specific proteins. Here, using ciprofloxacin as a DNA damaging agent specific to the organelles, we show that plastids and mitochondria can repair DNA double-strand breaks through an error-prone pathway similar to the microhomology-mediated break-induced replication observed in humans, yeast, and bacteria. This pathway is negatively regulated by the single-stranded DNA (ssDNA) binding proteins from the Whirly family, thus indicating that these proteins could contribute to the accurate repair of plant organelle genomes. To understand the role of Whirly proteins in this process, we solved the crystal structures of several Whirly-DNA complexes. These reveal a nonsequence-specific ssDNA binding mechanism in which DNA is stabilized between domains of adjacent subunits and rendered unavailable for duplex formation and/or protein interactions. Our results suggest a model in which the binding of Whirly proteins to ssDNA would favor accurate repair of DNA double-strand breaks over an error-prone microhomology-mediated break-induced replication repair pathway.