Summary information and primary citation
- PDB-id
- 1uhy; DSSR-derived features in text and JSON formats
- Class
- DNA
- Method
- X-ray (1.7 Å)
- Summary
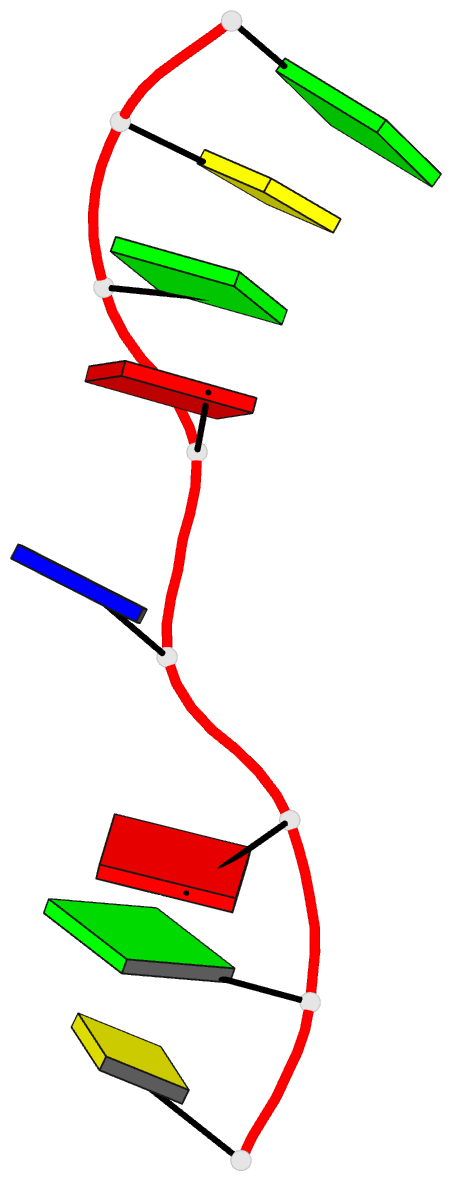
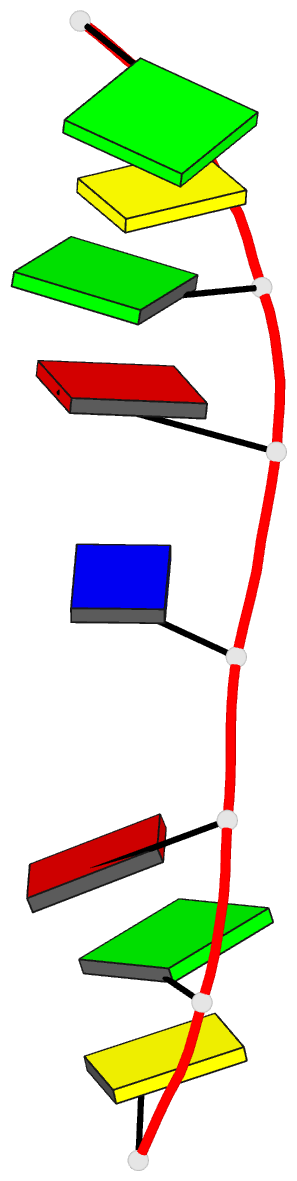
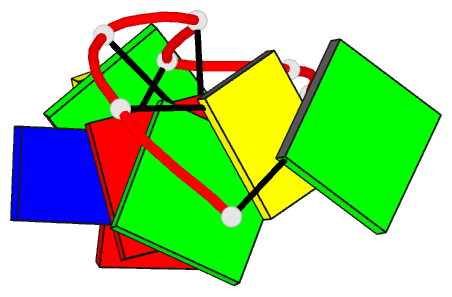
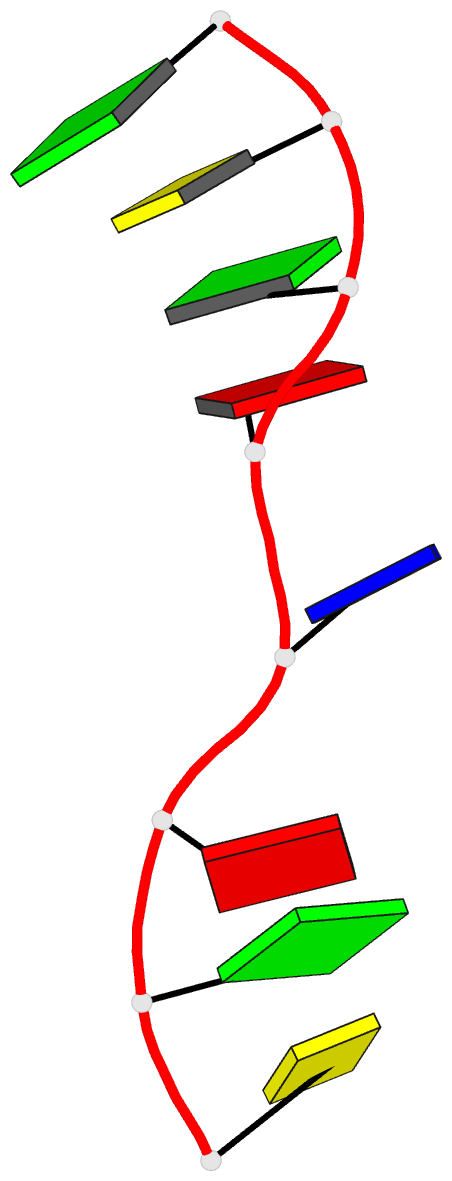
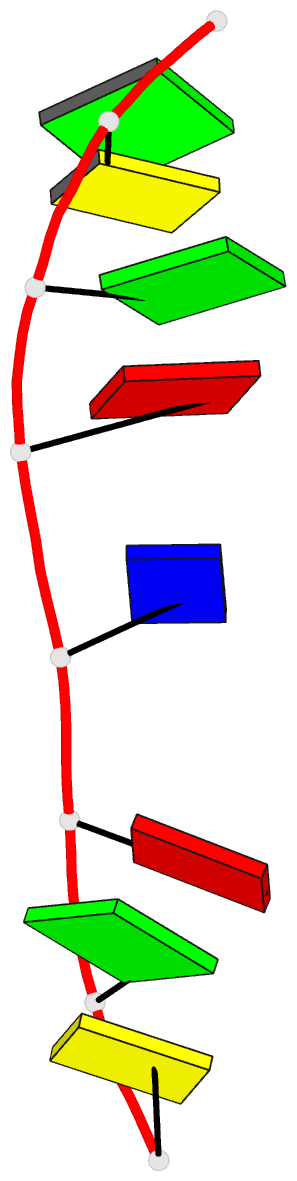
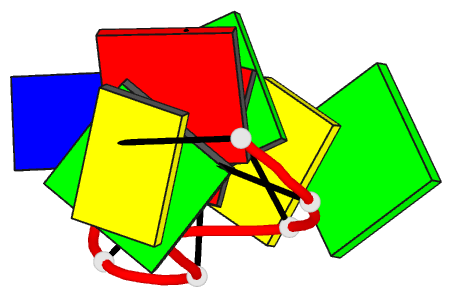
- Crystal structure of d(gcgatagc): the base-intercalated duplex
- Reference
- Kondo J, Umeda SI, Fujita K, Sunami T, Takenaka A (2004): "X-ray analyses of d(GCGAXAGC) containing G and T at X: the base-intercalated duplex is still stable even in point mutants at the fifth residue." J.Synchrotron Radiat., 11, 117-120. doi: 10.1107/S0909049503023562.
- Abstract
- DNA fragments containing the sequence d(GCGAAAGC) prefer to adopt a base-intercalated (zipper-like) duplex in the crystalline state. To investigate effects of point mutation at the 5th residue on the structure, two crystal structures of d(GCGAGAGC) and d(GCGATAGC) have been determined by X-ray crystallography. In the respective crystals, the two octamers related by a crystallographic two-fold symmetry are aligned in an anti-parallel fashion and associated to each other to form a duplex, suggesting that the base-intercalated duplex is stable even when the 5th residue is mutated with other bases. The sheared G3:A6 pair formation makes the two phosphate backbones closer and facilitates formation of the A-X*-X-A* base-intercalated motif. The three duplexes are assembled around the three-fold axis, and their 3rd and 4th residues are bound to the hexamine cobalt chloride. The central 5th residues are bound to another cation.