Summary information and primary citation
- PDB-id
-
1oct;
SNAP-derived features in text and
JSON formats
- Class
- transcription-DNA
- Method
- X-ray (3.0 Å)
- Summary
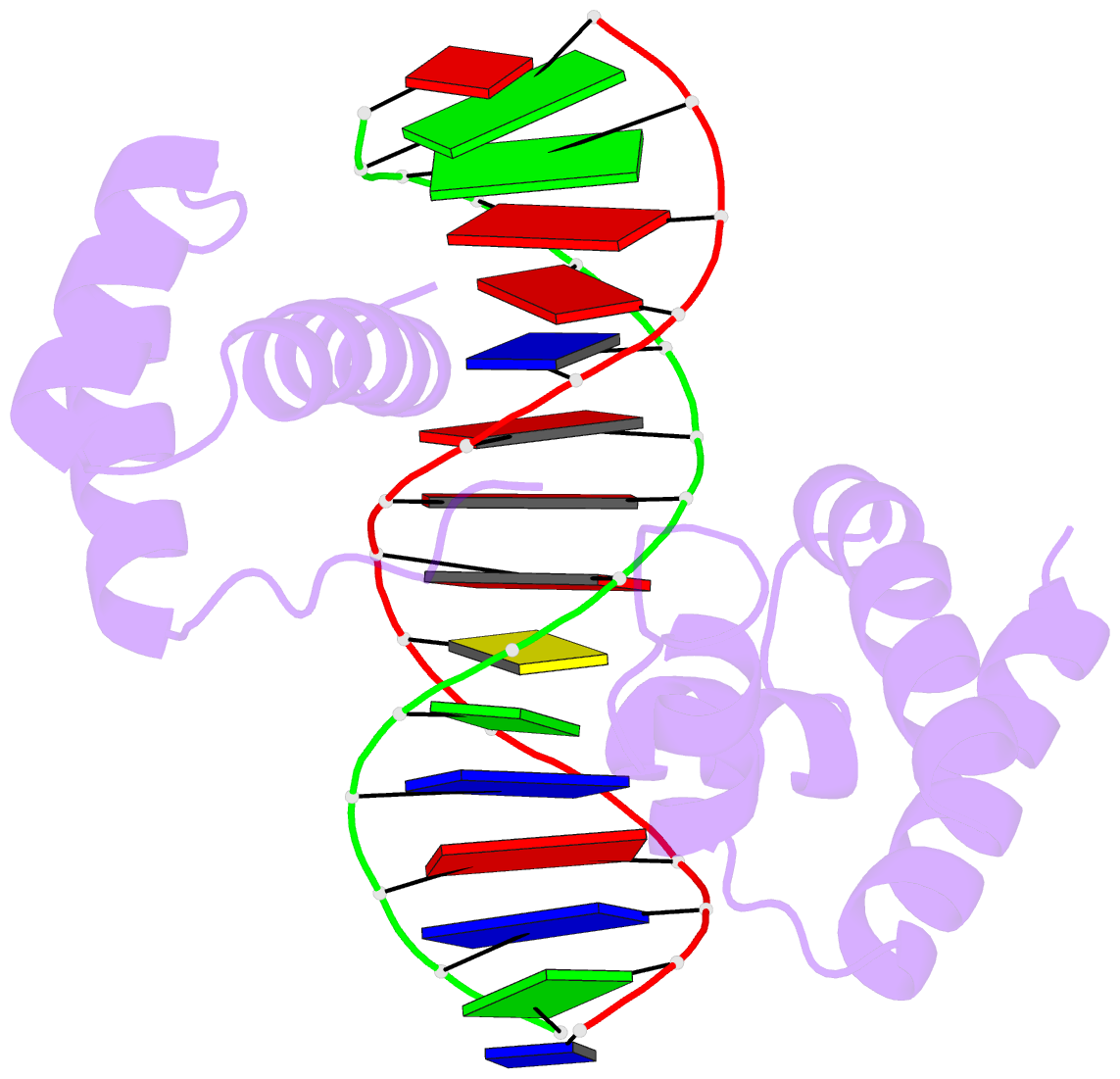
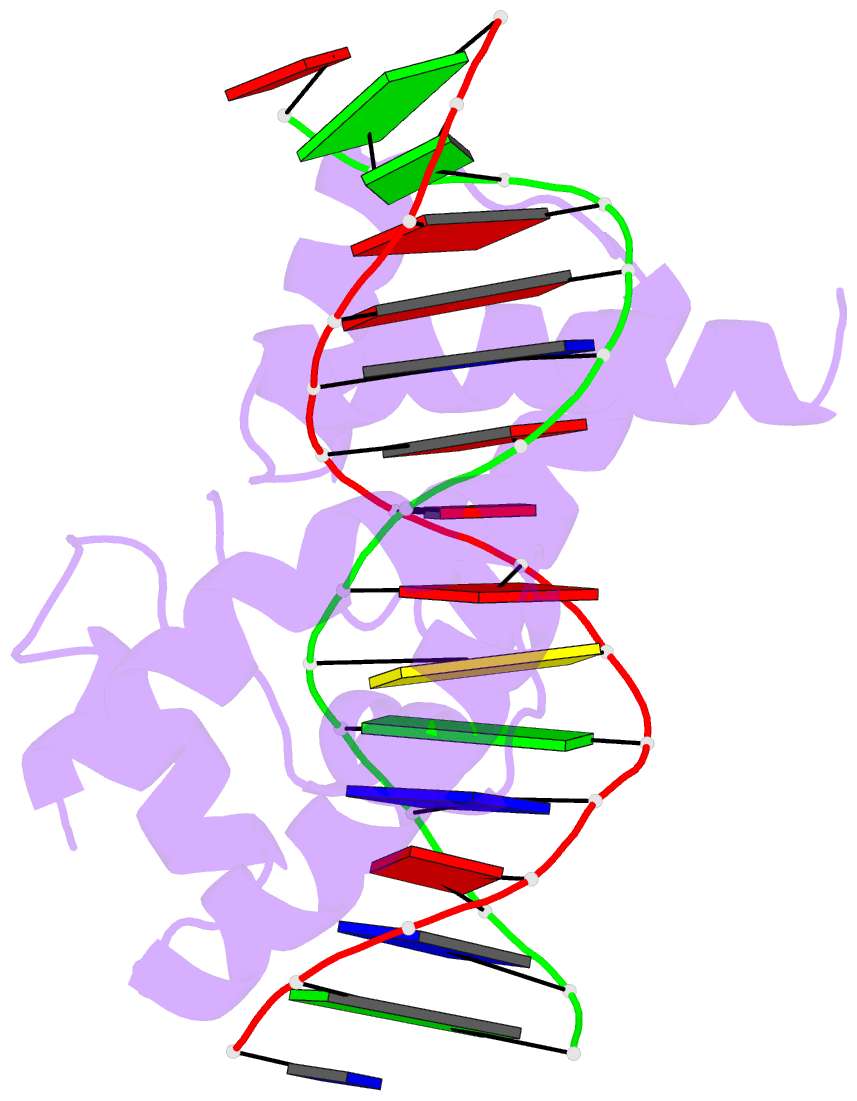
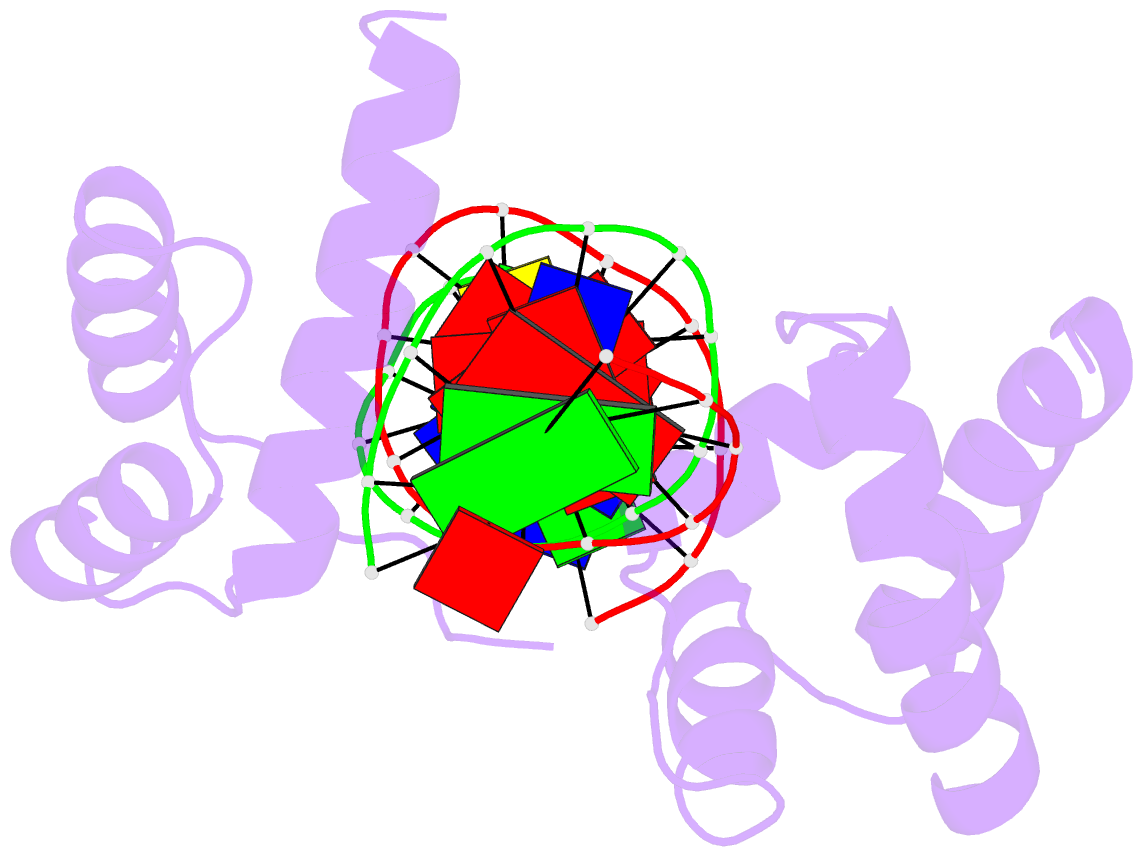
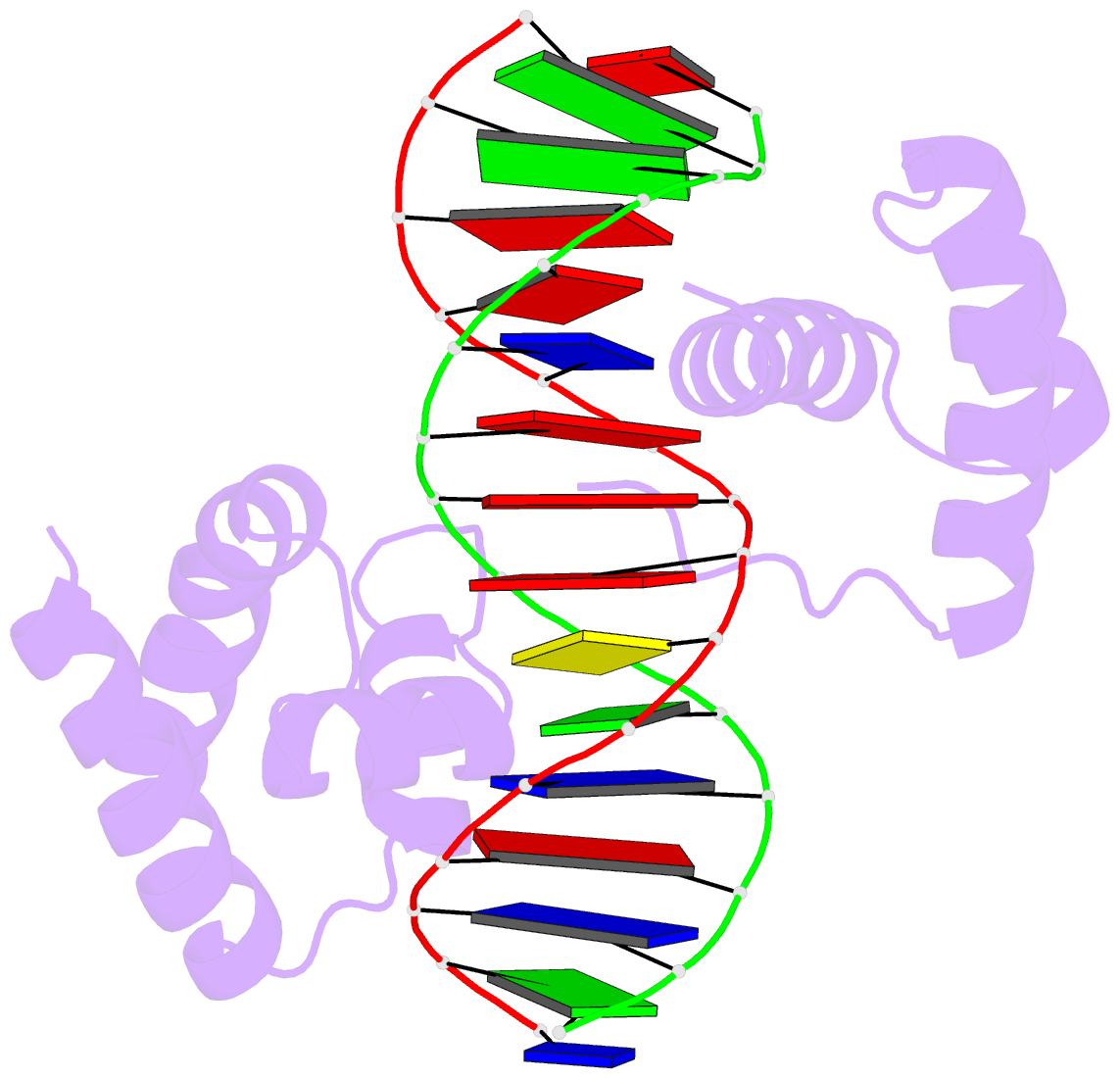
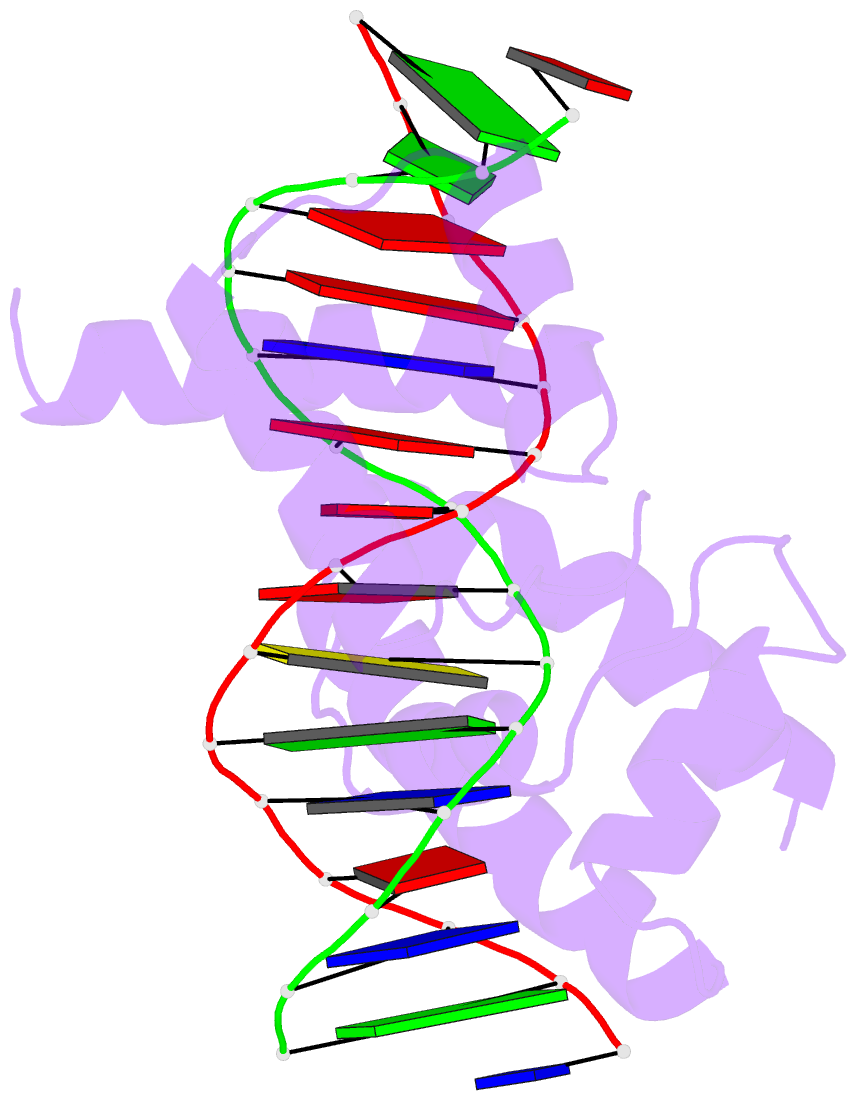
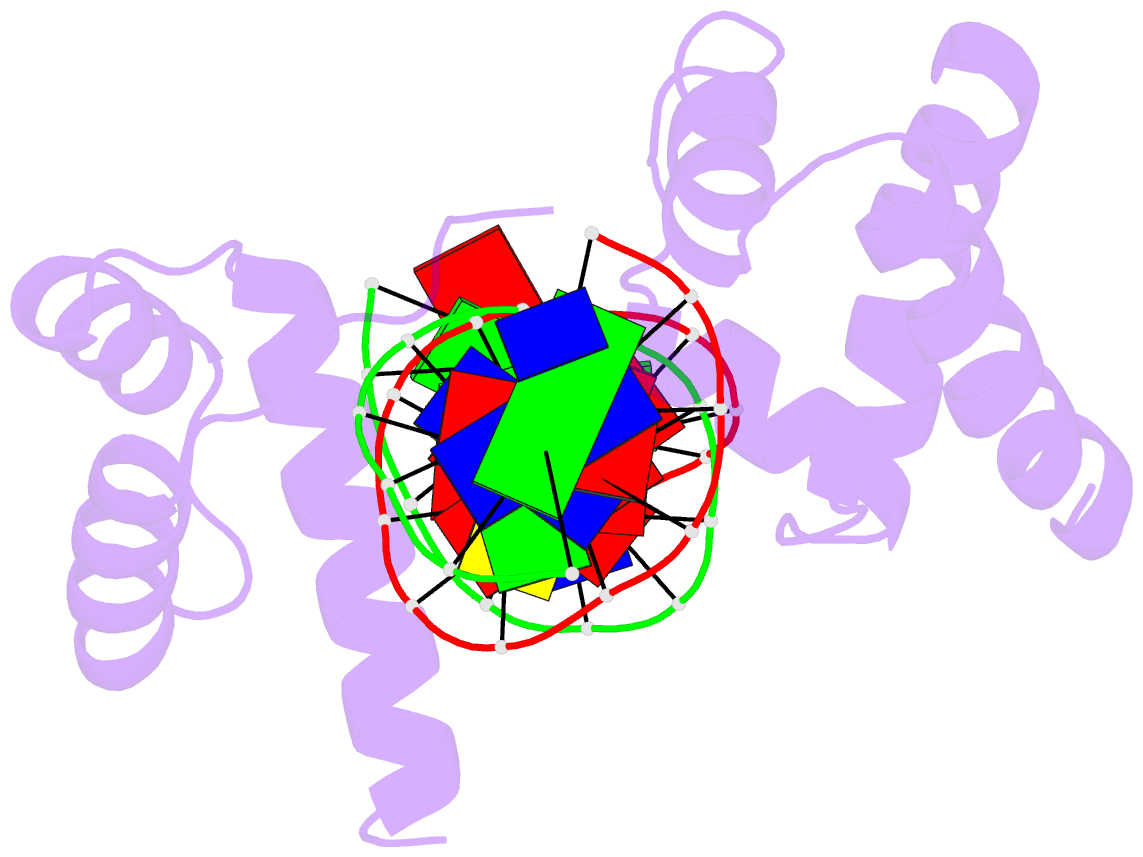
- Crystal structure of the oct-1 pou domain bound to an
octamer site: DNA recognition with tethered DNA-binding
modules
- Reference
-
Klemm JD, Rould MA, Aurora R, Herr W, Pabo CO (1994):
"Crystal
structure of the Oct-1 POU domain bound to an octamer
site: DNA recognition with tethered DNA-binding
modules." Cell(Cambridge,Mass.),
77, 21-32. doi: 10.1016/0092-8674(94)90231-3.
- Abstract
- The structure of an Oct-1 POU domain-octamer DNA
complex has been solved at 3.0 A resolution. The
POU-specific domain contacts the 5' half of this site
(ATGCAAAT), and as predicted from nuclear magnetic
resonance studies, the structure, docking, and contacts are
remarkably similar to those of the lambda and 434
repressors. The POU homeodomain contacts the 3' half of
this site (ATGCAAAT), and the docking is similar to that of
the engrailed, MAT alpha 2, and Antennapedia homeodomains.
The linker region is not visible and there are no
protein-protein contacts between the domains, but
overlapping phosphate contacts near the center of the
octamer site may favor cooperative binding. This novel
arrangement raises important questions about cooperativity
in protein-DNA recognition.