Summary information and primary citation
- PDB-id
- 1d54; DSSR-derived features in text and JSON formats
- Class
- DNA
- Method
- X-ray (1.4 Å)
- Summary
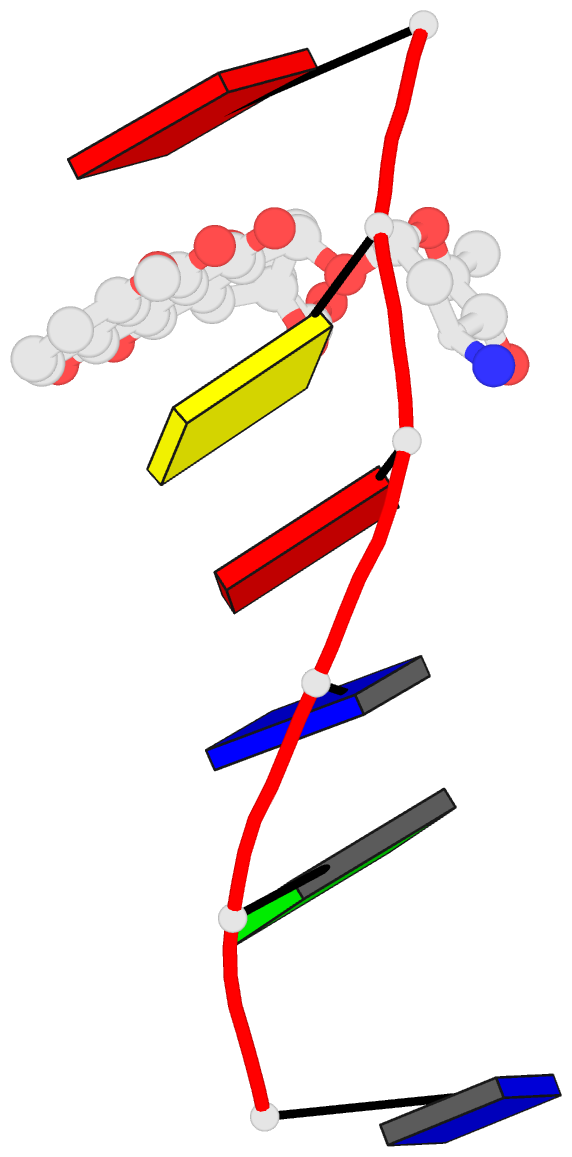
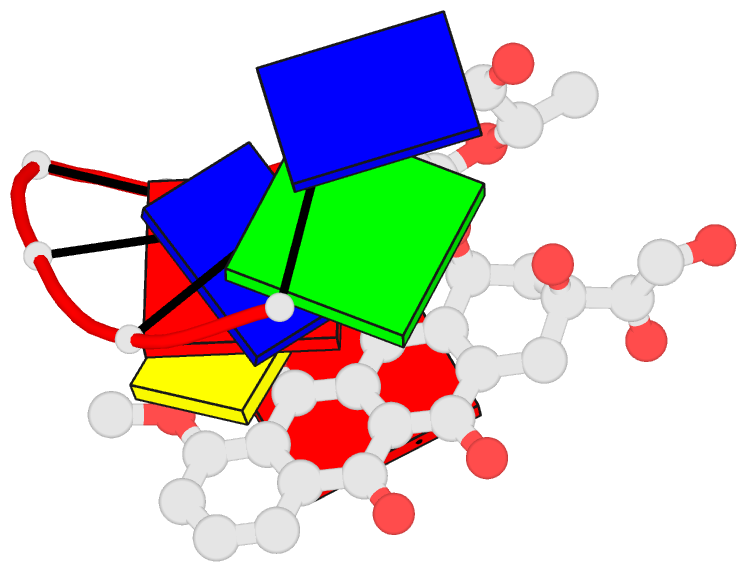
- Anthracycline binding to DNA: high resolution structure of d(tgtaca) complexed with 4'-epiadriamycin
- Reference
- Leonard GA, Brown T, Hunter WN (1992): "Anthracycline binding to DNA. High-resolution structure of d(TGTACA) complexed with 4'-epiadriamycin." Eur.J.Biochem., 204, 69-74. doi: 10.1111/j.1432-1033.1992.tb16606.x.
- Abstract
- Crystallographic methods have been applied to determine the high-resolution structure of the complex formed between the self-complementary oligonucleotide d(TGTACA) and the anthracycline antibiotic 4'-epiadriamycin. The complex crystallises in the tetragonal system, space group P4(1)2(1)2 with a = 2.802 nm and c = 5.293 nm, and an asymmetric unit consisting of a single DNA strand, one drug molecule and 34 solvent molecules. The refinement converged with an R factor of 0.17 for the 2381 reflections with F greater than or equal to 3 sigma F in the resolution range 0.70-0.14 nm. Two asymmetric units associate such that a distorted B-DNA-type hexanucleotide duplex is formed incorporating two drug molecules that are intercalated at the TpG steps. The amino sugar of 4'-epiadriamycin binds in the minor groove of the duplex and displays different interactions from those observed in previously determined structures. Interactions between the hydrophilic groups of the amino sugar and the oligonucleotide are all mediated by solvent molecules. Ultraviolet melting measurements and comparison with other anthracycline-DNA complexes suggest that these indirect interactions have a powerful stabilising effect on the complex.