Summary information and primary citation
- PDB-id
- 1byx; DSSR-derived features in text and JSON formats
- Class
- DNA-RNA hybrid
- Method
- NMR
- Summary
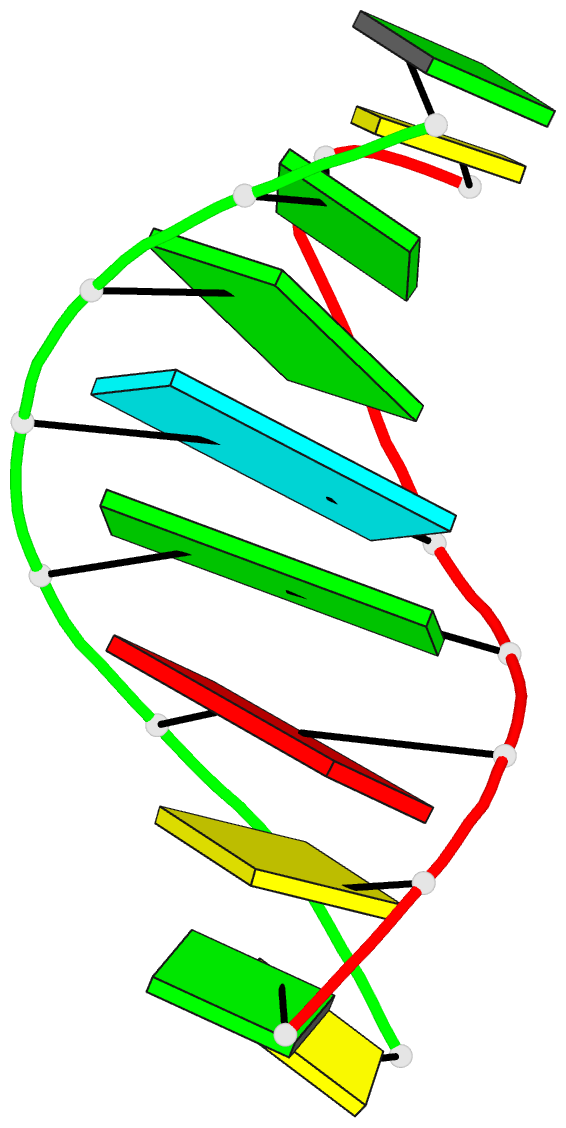
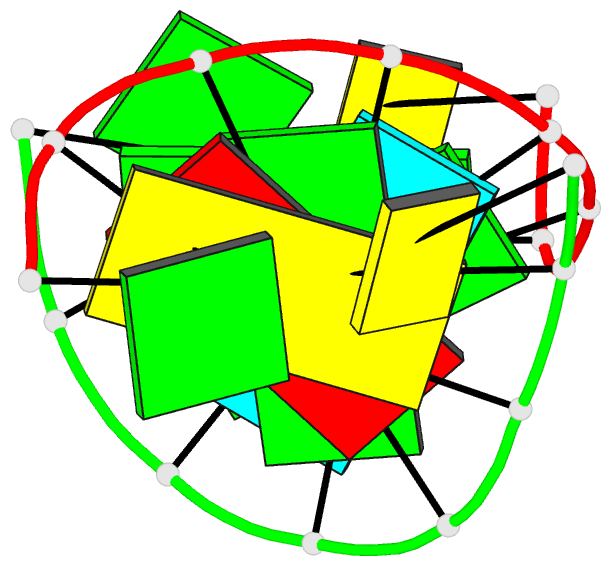
- Chimeric hybrid duplex r(gcaguggc).r(gcca)d(ctgc) comprising the trna-DNA junction formed during initiation of hiv-1 reverse transcription
- Reference
- Szyperski T, Gotte M, Billeter M, Perola E, Cellai L, Heumann H, Wuthrich K (1999): "NMR structure of the chimeric hybrid duplex r(gcaguggc).r(gcca)d(CTGC) comprising the tRNA-DNA junction formed during initiation of HIV-1 reverse transcription." J.Biomol.NMR, 13, 343-355. doi: 10.1023/A:1008350604637.
- Abstract
- A high-quality NMR solution structure of the chimeric hybrid duplex r(gcaguggc).r(gcca)d(CTGC) was determined using the program DYANA with its recently implemented new module FOUND, which performs exhaustive conformational grid searches for dinucleotides. To ensure conservative data interpretation, the use of 1H-1H lower distance limit constraints was avoided. The duplex comprises the tRNA-DNA junction formed during the initiation of HIV-1 reverse transcription. It forms an A-type double helix that exhibits distinct structural deviations from a standard A-conformation. In particular, the minor groove is remarkably narrow, and its width decreases from about 7.5 A in the RNA/RNA stem to about 4.5 A in the RNA/DNA segment. This is unexpected, since minor groove widths for A-RNA and RNA/DNA hybrid duplexes of approximately 11 A and approximately 8.5 A, respectively, were previously reported. The present, new structure supports that reverse transcriptase-associated RNaseH specificity is related primarily to conformational adaptability of the nucleic acid in 'induced-fit'-type interactions, rather than the minor groove width of a predominantly static nucleic acid duplex.